Orbit stability during van der Meer scan
The goal of this presentation is to share a (preliminary) analysis of the orbit stability of one of the 2018 van der Meer scan (FILL 6868).
From the analysis carried out from the experiments, there are evidences of hysteresis effects during the scan.
I fact, the scan consists in varying, using special orbit bumps, the beams positions at the IP (namely the H/V separation of the two beams). To do so the current of special orbit correctors is varied and this can potentially produce hysteretic behaviour of the magnets themselves.
In the following we will show the approach we used to verify and assess the hysteretics behaviour of the magnets.
Analysis can be run from /eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/OrbitStabilityVdM.ipynb (stack 95py3).
 From https://i.redd.it/c2xhv8qhn0h01.jpg
From https://i.redd.it/c2xhv8qhn0h01.jpg
# Using the stack 95py3
mySource='/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/OrbitStabilityVdM.ipynb'
import cl2pd # do "pip install --user git+https://github.com/sterbini/cl2pd.git@py3" to install
from cl2pd import importData
from cl2pd import plotFunctions
from cl2pd import dotdict
from cl2pd import bbFunctions
from matplotlib import dates as dates
dotdict=dotdict.dotdict
pd=importData.pd # is the pandas package
np=importData.np # is the numpy package
cals=importData.cals # pytimber log class
import matplotlib.pyplot as plt
get_ipython().magic('matplotlib inline')
%config InlineBackend.figure_format = 'retina' # retina display
myTitle='FILL 6868, 30th June 2018'
myFill=6868
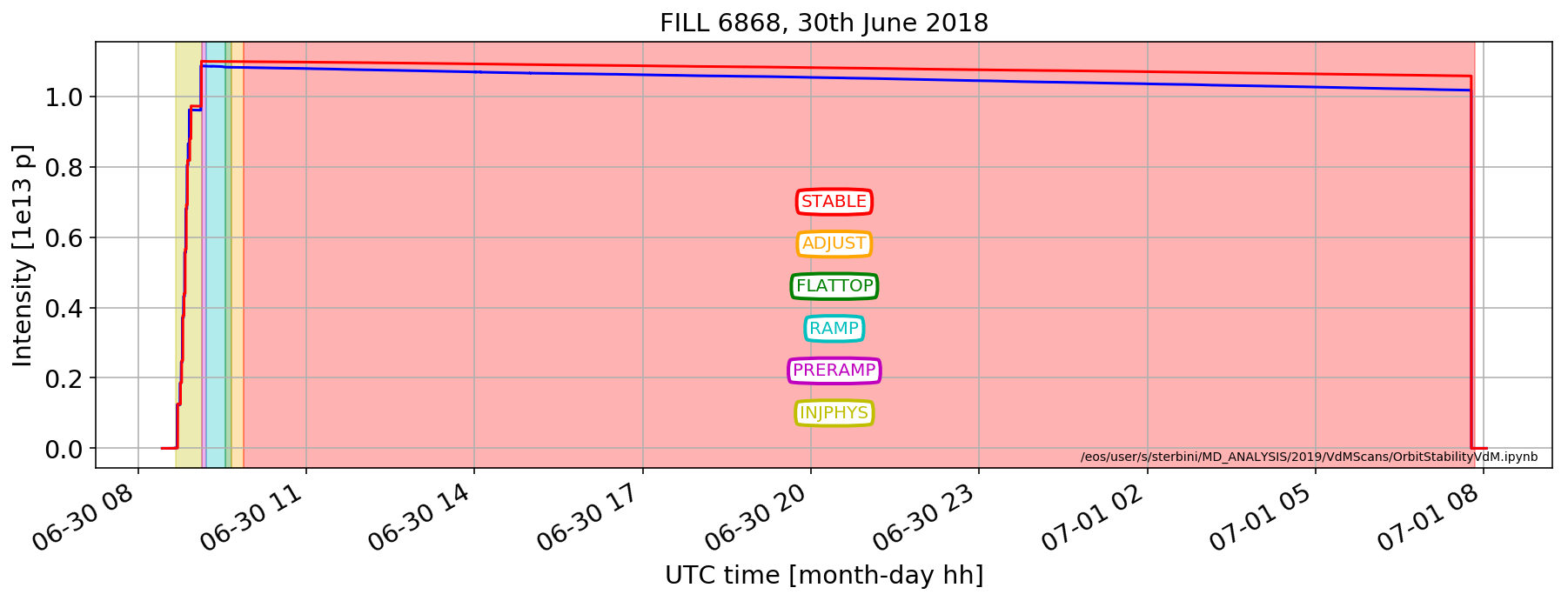
Fill 6868:
This is the usual cycle overview.
# Analysis of the fill
fillDF=importData.LHCFillsByNumber([myFill])
fillDF[fillDF['mode']=='FILL']
| mode | startTime | endTime | duration | |
|---|---|---|---|---|
| 6868 | FILL | 2018-06-30 08:25:45.239000082+00:00 | 2018-07-01 08:02:33.473999977+00:00 | 23:36:48.234999 |
params = {'legend.fontsize': 'x-large',
'figure.figsize': (15, 5),
'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large'}
plt.rcParams.update(params)
# use the "ignore" warning with care
import warnings
warnings.filterwarnings("ignore")
plotFunctions.plotLHCFill(fillDF,myTitle,pd.Timestamp('2018-06-30 08:25:45.239000082'))
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')

A very long fill: protons travelled more than Voyager 1, the farthest man-crafted object (see https://voyager.jpl.nasa.gov/).
Beam intensities
myDF=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B%:BEAM_INTENSITY'],[myFill])
myFigIntensityDecay=plt.figure()
plt.plot(myDF['LHC.BCTFR.A6R4.B1:BEAM_INTENSITY'].dropna()/1e13, color='b')
(myDF['LHC.BCTFR.A6R4.B2:BEAM_INTENSITY'].dropna()/1e13).plot( color='r')
plt.grid('on')
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('Intensity [1e13 p]');
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='INJPHYS'], color='y',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='PRERAMP'], color='m',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='RAMP'], color='c',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='FLATTOP'], color='g',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='ADJUST'], color='orange',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='STABLE'], color='r',alpha=.3)
t1=dates.date2num(pd.Timestamp('2018-06-30 20:25'))
if 1:
for i,j,k in zip(['INJPHYS','PRERAMP','RAMP','FLATTOP','ADJUST','STABLE'],np.arange(0,6),
['y','m','c','g','orange','r']):
plotFunctions.setArrowLabel(plt.gca(), label=i,
arrowPosition=(t1, .1+j*.12),
labelPosition=(t1, .1+j*.12),
myColor=k, arrowArc_rad=-0.2);
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
plt.title(myTitle);
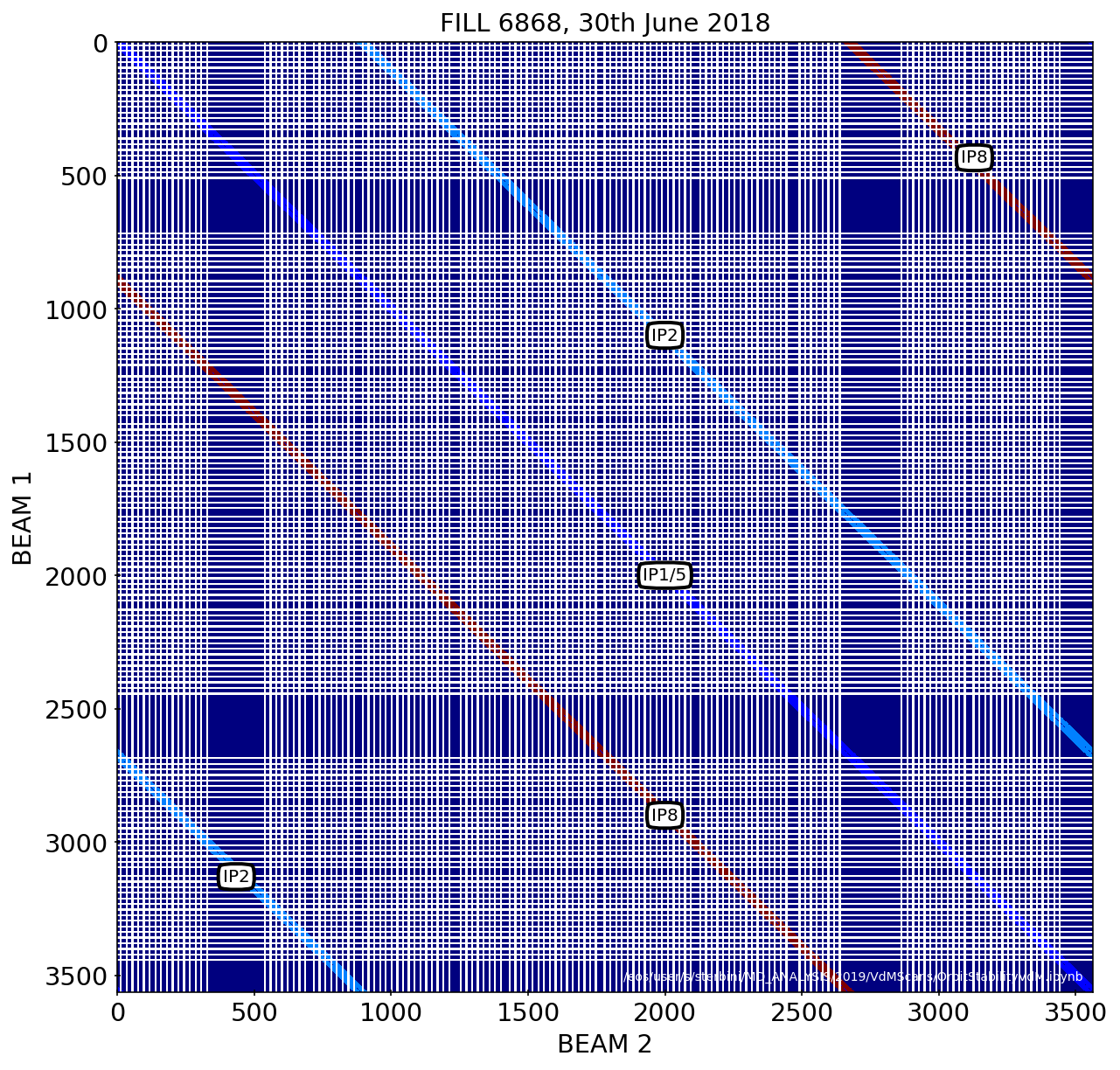
Filling scheme
# BB Matrix, considering 25 encouters
nbLR = 25
fillingSchemeDF=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B%:BUNCH_FILL_PATTERN'], myFill, 'RAMP', flag='last')
print(f"Number of bunches in B1: \
{len(np.where(fillingSchemeDF['LHC.BCTFR.A6R4.B1:BUNCH_FILL_PATTERN'].iloc[0])[0])}")
print(f"Number of bunches in B2: \
{len(np.where(fillingSchemeDF['LHC.BCTFR.A6R4.B1:BUNCH_FILL_PATTERN'].iloc[0])[0])}")
Number of bunches in B1: 140
Number of bunches in B2: 140
BBMatrixLHC=bbFunctions.computeBBMatrix(numberOfLRToConsider=nbLR)
# We plot it
B1_bunches=np.where(fillingSchemeDF.iloc[0]['LHC.BCTFR.A6R4.B1:BUNCH_FILL_PATTERN'])[0]
B2_bunches=np.where(fillingSchemeDF.iloc[0]['LHC.BCTFR.A6R4.B2:BUNCH_FILL_PATTERN'])[0]
plotFunctions.plot_BBMatrix(BBMatrixLHC, B1_bunches, B2_bunches,alpha=1,width=1)
plt.title(myTitle);
plotFunctions.setSourcePlot(plt.gca(),mySource,color='w')
B1CollisionSchedule=bbFunctions.B1CollisionScheduleDF(
fillingSchemeDF['LHC.BCTFR.A6R4.B1:BUNCH_FILL_PATTERN'].iloc[0],
fillingSchemeDF['LHC.BCTFR.A6R4.B2:BUNCH_FILL_PATTERN'].iloc[0],
16,
)
B1CollisionSchedule.head(2)
| HO partner in ALICE | # of LR in ALICE | BB partners in ALICE | Positions in ALICE | HO partner in ATLAS/CMS | # of LR in ATLAS/CMS | BB partners in ATLAS/CMS | Positions in ATLAS/CMS | HO partner in LHCB | # of LR in LHCB | BB partners in LHCB | Positions in LHCB | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 12 | NaN | 2 | [896.0, 917.0] | [-7.0, 14.0] | 12.0 | 0 | [12.0] | [0.0] | NaN | 0 | [] | [] |
| 33 | NaN | 2 | [917.0, 938.0] | [-7.0, 14.0] | 33.0 | 0 | [33.0] | [0.0] | NaN | 0 | [] | [] |
B2CollisionSchedule=bbFunctions.B2CollisionScheduleDF(
fillingSchemeDF['LHC.BCTFR.A6R4.B1:BUNCH_FILL_PATTERN'].iloc[0],
fillingSchemeDF['LHC.BCTFR.A6R4.B2:BUNCH_FILL_PATTERN'].iloc[0],
16,
)
B1CollisionSchedule.head(2)
| HO partner in ALICE | # of LR in ALICE | BB partners in ALICE | Positions in ALICE | HO partner in ATLAS/CMS | # of LR in ATLAS/CMS | BB partners in ATLAS/CMS | Positions in ATLAS/CMS | HO partner in LHCB | # of LR in LHCB | BB partners in LHCB | Positions in LHCB | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 12 | NaN | 2 | [896.0, 917.0] | [-7.0, 14.0] | 12.0 | 0 | [12.0] | [0.0] | NaN | 0 | [] | [] |
| 33 | NaN | 2 | [917.0, 938.0] | [-7.0, 14.0] | 33.0 | 0 | [33.0] | [0.0] | NaN | 0 | [] | [] |
print(f"Collision in ALICE: {len(B1CollisionSchedule['HO partner in ALICE'].dropna())}")
print(f"Collision in ATLAS/CMS: {len(B1CollisionSchedule['HO partner in ATLAS/CMS'].dropna())}")
print(f"Collision in LHCB: {len(B1CollisionSchedule['HO partner in LHCB'].dropna())}")
Collision in ALICE: 32
Collision in ATLAS/CMS: 124
Collision in LHCB: 23
# Selecting the B1 bunches
# 1. colliding HO in ATLAS and CMS
# 2. without HO collision in ALICE
# 3. without HO collision and LHCB
# 4. without BBLR in ALICE
B1CollisionSchedule[(~np.isnan(B1CollisionSchedule['HO partner in ATLAS/CMS'])) & \
(np.isnan(B1CollisionSchedule['HO partner in ALICE'])) & \
(np.isnan(B1CollisionSchedule['HO partner in LHCB'])) & \
(B1CollisionSchedule['# of LR in ALICE']==0)
]
| HO partner in ALICE | # of LR in ALICE | BB partners in ALICE | Positions in ALICE | HO partner in ATLAS/CMS | # of LR in ATLAS/CMS | BB partners in ATLAS/CMS | Positions in ATLAS/CMS | HO partner in LHCB | # of LR in LHCB | BB partners in LHCB | Positions in LHCB | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1779 | NaN | 0 | [] | [] | 1779.0 | 0 | [1779.0] | [0.0] | NaN | 1 | [896.0] | [11.0] |
| 1800 | NaN | 0 | [] | [] | 1800.0 | 0 | [1800.0] | [0.0] | NaN | 2 | [896.0, 917.0] | [-10.0, 11.0] |
| 1821 | NaN | 0 | [] | [] | 1821.0 | 0 | [1821.0] | [0.0] | NaN | 2 | [917.0, 938.0] | [-10.0, 11.0] |
| 1842 | NaN | 0 | [] | [] | 1842.0 | 0 | [1842.0] | [0.0] | NaN | 2 | [938.0, 959.0] | [-10.0, 11.0] |
| 1863 | NaN | 0 | [] | [] | 1863.0 | 0 | [1863.0] | [0.0] | NaN | 2 | [959.0, 980.0] | [-10.0, 11.0] |
| 1884 | NaN | 0 | [] | [] | 1884.0 | 0 | [1884.0] | [0.0] | NaN | 2 | [980.0, 1001.0] | [-10.0, 11.0] |
| 1905 | NaN | 0 | [] | [] | 1905.0 | 0 | [1905.0] | [0.0] | NaN | 2 | [1001.0, 1022.0] | [-10.0, 11.0] |
| 1926 | NaN | 0 | [] | [] | 1926.0 | 0 | [1926.0] | [0.0] | NaN | 2 | [1022.0, 1043.0] | [-10.0, 11.0] |
| 1947 | NaN | 0 | [] | [] | 1947.0 | 0 | [1947.0] | [0.0] | NaN | 2 | [1043.0, 1064.0] | [-10.0, 11.0] |
| 3032 | NaN | 0 | [] | [] | 3032.0 | 0 | [3032.0] | [0.0] | NaN | 2 | [2128.0, 2149.0] | [-10.0, 11.0] |
| 3053 | NaN | 0 | [] | [] | 3053.0 | 0 | [3053.0] | [0.0] | NaN | 2 | [2149.0, 2170.0] | [-10.0, 11.0] |
| 3074 | NaN | 0 | [] | [] | 3074.0 | 0 | [3074.0] | [0.0] | NaN | 2 | [2170.0, 2191.0] | [-10.0, 11.0] |
| 3095 | NaN | 0 | [] | [] | 3095.0 | 0 | [3095.0] | [0.0] | NaN | 2 | [2191.0, 2212.0] | [-10.0, 11.0] |
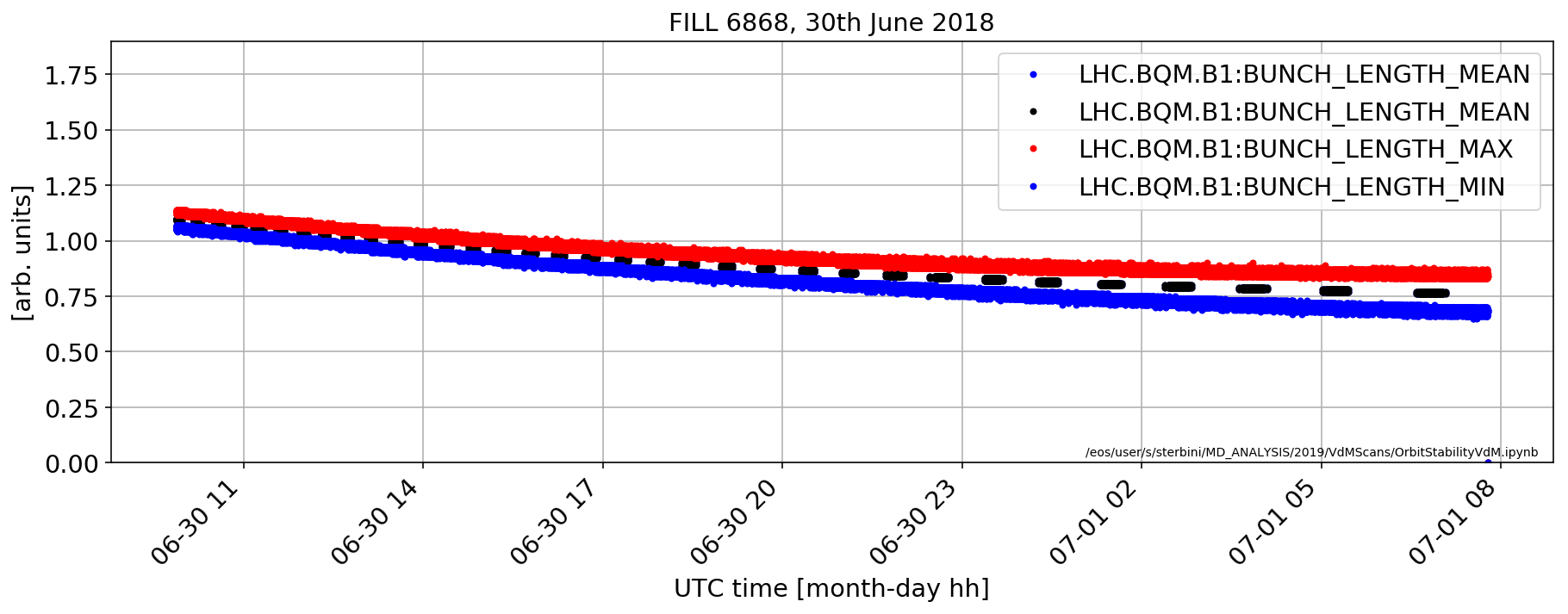
Longitudinal sigma
if 1:
rawDF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BQM.pickle')
else:
rawDF=importData.LHCCals2pd(['LHC.BQM.B%:BUNCH_LENGTH_M%'],[myFill],'STABLE')
rawDF.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BQM.pickle')
plt.plot(rawDF['LHC.BQM.B1:BUNCH_LENGTH_MEAN'].dropna()*1e9,'b.')
(rawDF['LHC.BQM.B1:BUNCH_LENGTH_MEAN'].dropna()*1e9).plot(marker='.',color='k',ls='none')
(rawDF['LHC.BQM.B1:BUNCH_LENGTH_MAX'].dropna()*1e9).plot(marker='.',color='r',ls='none')
(rawDF['LHC.BQM.B1:BUNCH_LENGTH_MIN'].dropna()*1e9).plot(marker='.',color='b',ls='none')
plt.xlabel('time [hh:mm]')
plt.ylabel('mean bunch length (4$\sigma$) [ns]')
plt.grid('on')
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[arb. units]');
plt.legend(loc=1)
plt.ylim(0,1.9)
plt.xticks(rotation=45)
if 0:
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='INJPHYS'], color='y',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='PRERAMP'], color='m',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='RAMP'], color='c',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='FLATTOP'], color='g',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='SQUEEZE'], color='orange',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='ADJUST'], color='r',alpha=.3)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
plt.title(myTitle);
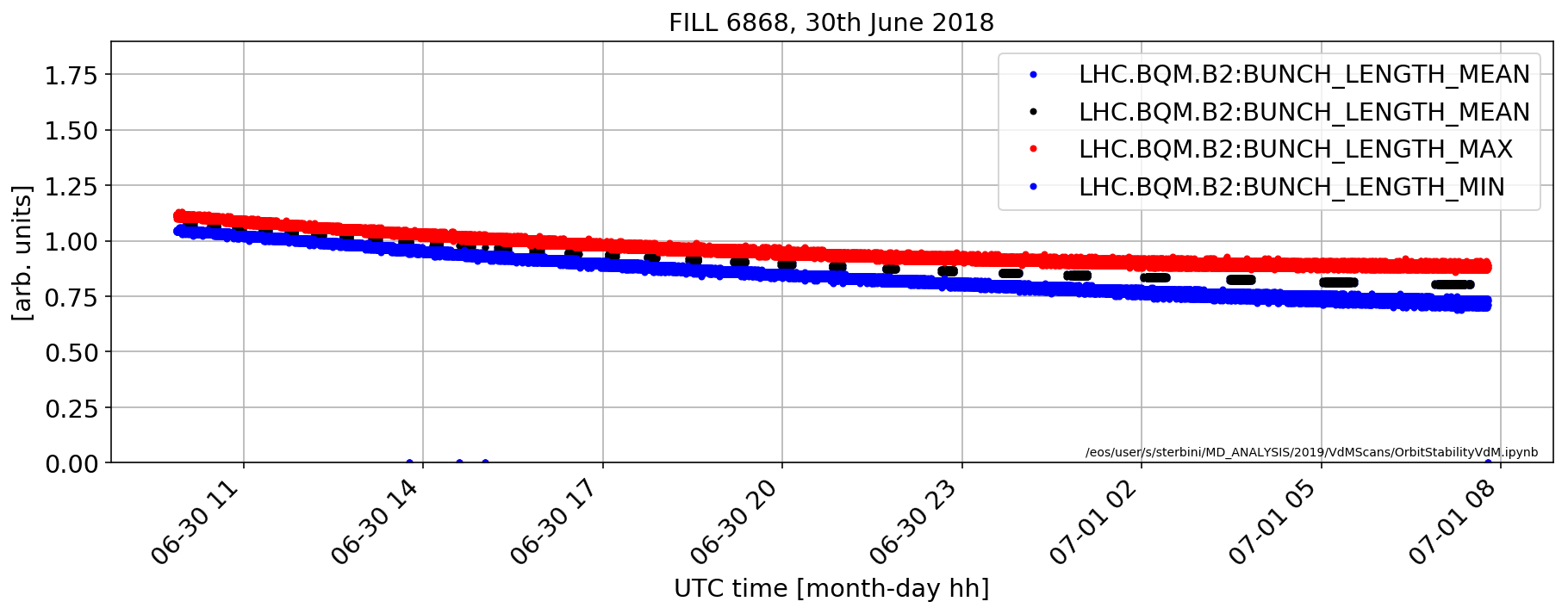
plt.plot(rawDF['LHC.BQM.B2:BUNCH_LENGTH_MEAN'].dropna()*1e9,'b.')
(rawDF['LHC.BQM.B2:BUNCH_LENGTH_MEAN'].dropna()*1e9).plot(marker='.',color='k',ls='none')
(rawDF['LHC.BQM.B2:BUNCH_LENGTH_MAX'].dropna()*1e9).plot(marker='.',color='r',ls='none')
(rawDF['LHC.BQM.B2:BUNCH_LENGTH_MIN'].dropna()*1e9).plot(marker='.',color='b',ls='none')
plt.xlabel('time [hh:mm]')
plt.ylabel('mean bunch length (4$\sigma$) [ns]')
plt.grid('on')
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[arb. units]');
plt.legend(loc=1)
plt.ylim(0,1.9)
plt.xticks(rotation=45)
if 0:
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='INJPHYS'], color='y',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='PRERAMP'], color='m',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='RAMP'], color='c',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='FLATTOP'], color='g',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='SQUEEZE'], color='orange',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='ADJUST'], color='r',alpha=.3)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
plt.title(myTitle);
Intensity decay
if 1:
myDF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BCTFR.pickle')
else:
myDF=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B%:BUNCH_INTENSITY'],[myFill],['STABLE'])
myDF.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BCTFR.pickle')
for i in np.where(np.array(myDF['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'].iloc[0])>.2e11)[0]:
plt.plot(myDF['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'].apply(lambda x:x[i]))
plt.grid(True)
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[ppb]');
plt.xticks(rotation=45);
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
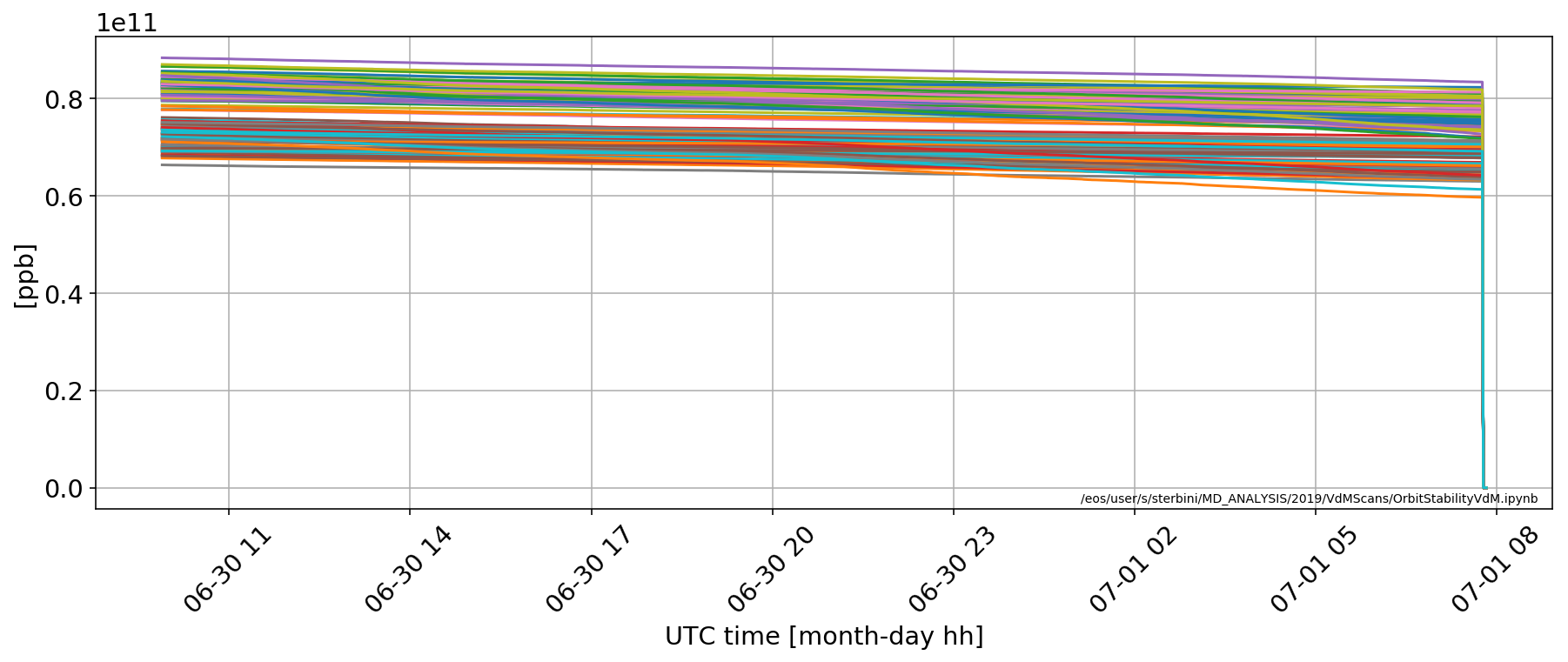
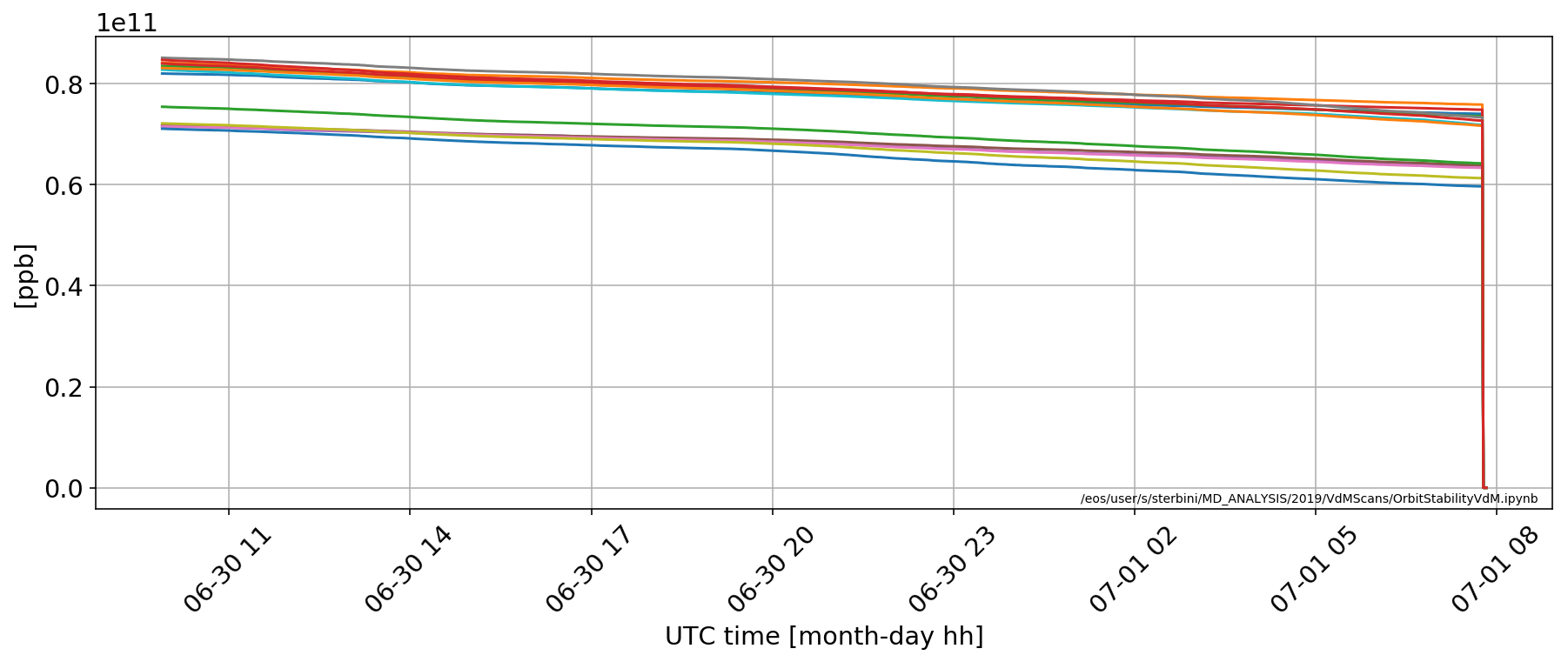
The problematic bunches are [1600, 1642, 1684, 1726, 1747, 3148, 3190, 3211, 3232, 3253, 3274, 3295, 3316, 3337] in B1.
for i in [1600, 1642, 1684, 1726, 1747, 3148, 3190, 3211, 3232, 3253, 3274, 3295, 3316, 3337]:
plt.plot(myDF['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'].apply(lambda x:x[i]))
plt.grid(True)
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[ppb]');
plt.xticks(rotation=45);
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
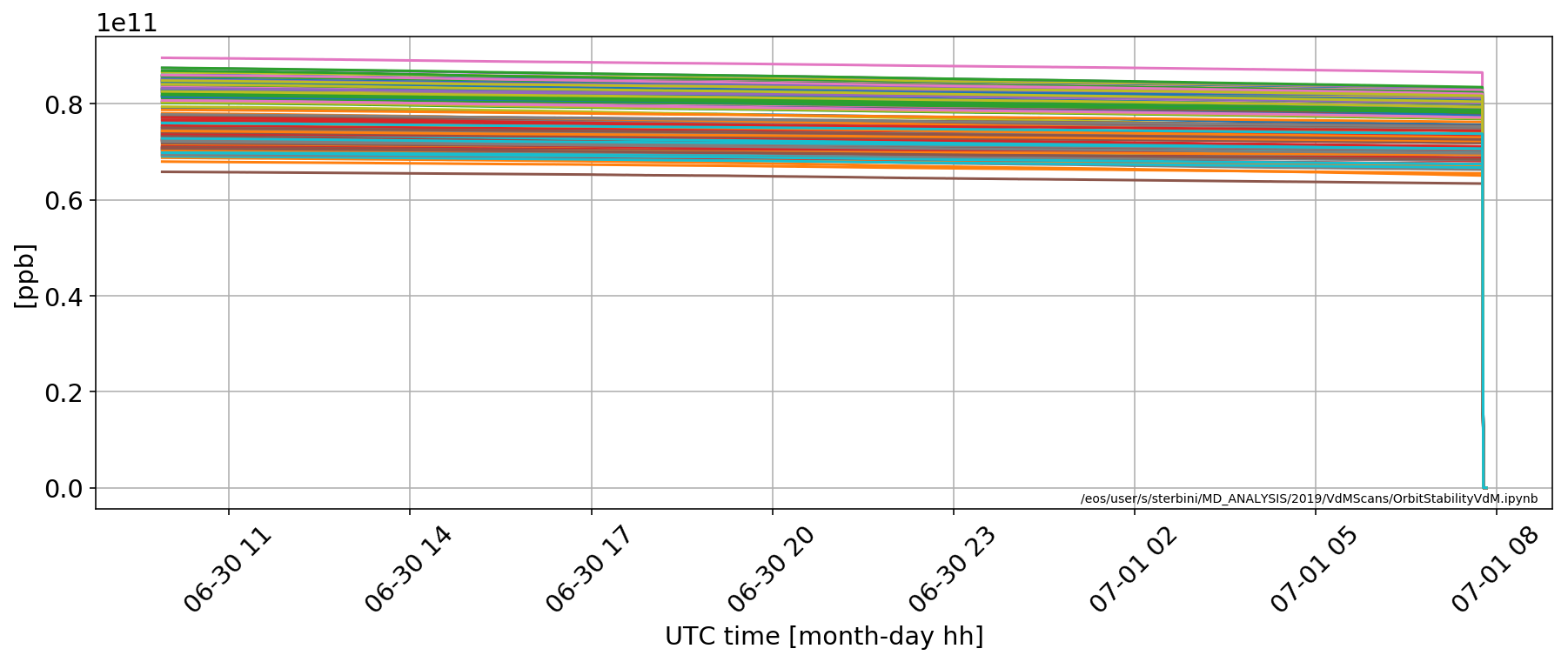
for i in np.where(np.array(myDF['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'].iloc[0])>.2e11)[0]:
plt.plot(myDF['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'].apply(lambda x:x[i]))
plt.grid('on')
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[ppb]');
plt.xticks(rotation=45);
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
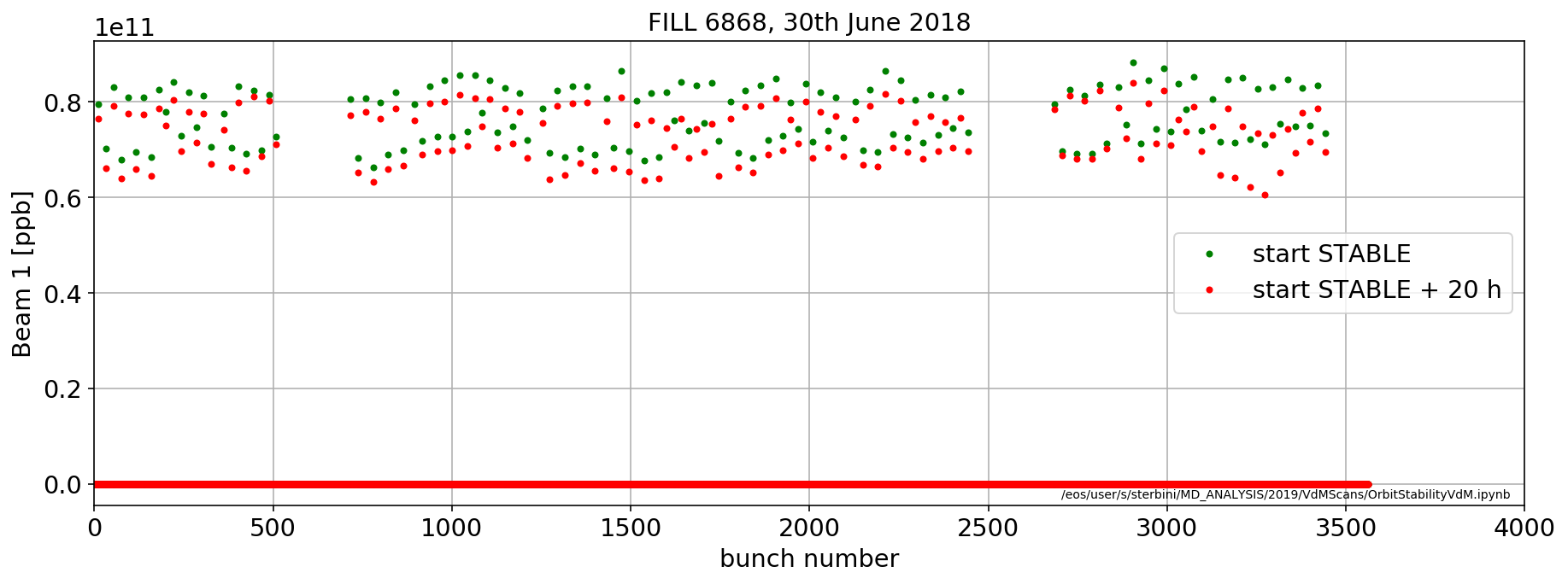
aux=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'],[myFill],['STABLE'],flag='next',offset=pd.Timedelta('0m'))
plt.plot(aux['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'].iloc[0],'.g',label='start STABLE')
aux=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'],[myFill],['STABLE'],flag='next',offset=pd.Timedelta('1200m'))
plt.plot(aux['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'].iloc[0],'.r',label='start STABLE + 20 h')
plt.xlim([0,4000])
plt.grid('on')
plt.xlabel('bunch number');
plt.ylabel('Beam 1 [ppb]');
plt.legend(loc='best')
plt.title(myTitle)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
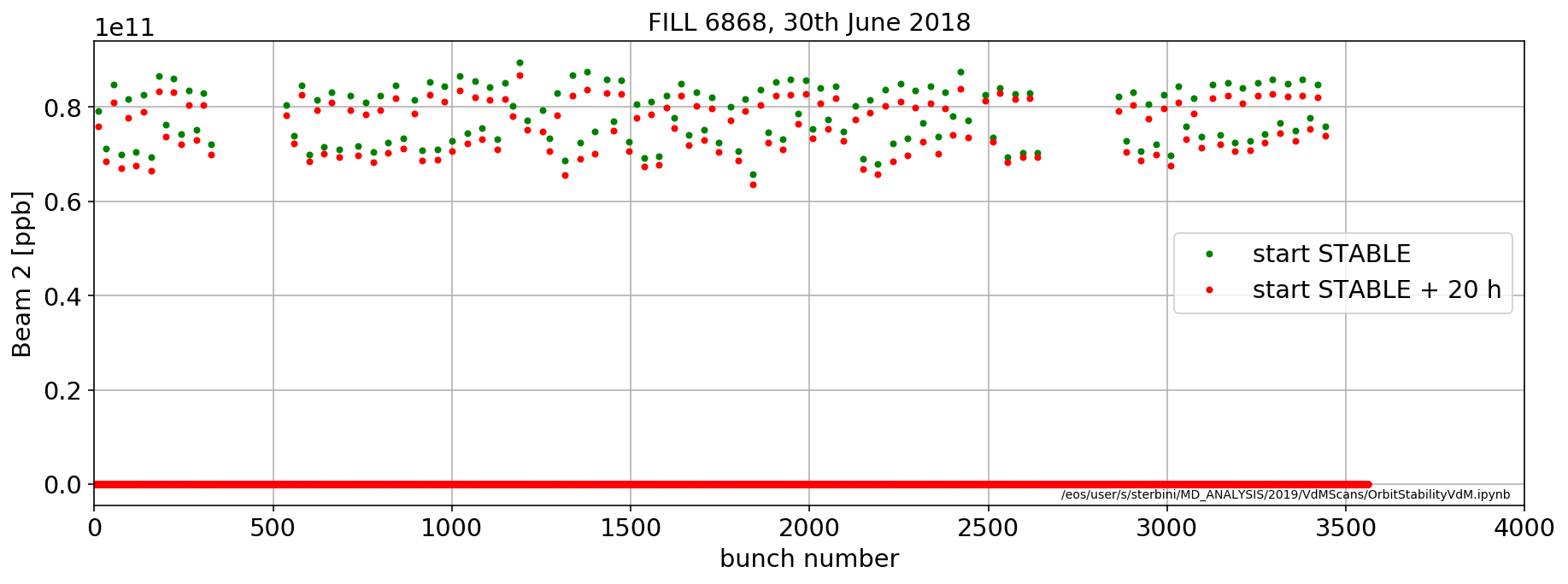
aux=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'],[myFill],['STABLE'],flag='next',offset=pd.Timedelta('0m'))
plt.plot(aux['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'].iloc[0],'.g',label='start STABLE')
aux=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'],[myFill],['STABLE'],flag='next',offset=pd.Timedelta('1200m'))
plt.plot(aux['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'].iloc[0],'.r',label='start STABLE + 20 h')
plt.xlim([0,4000])
plt.grid('on')
plt.xlabel('bunch number');
plt.ylabel('Beam 2 [ppb]');
plt.legend(loc='best')
plt.title(myTitle)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
plt.figure(figsize=(15,5))
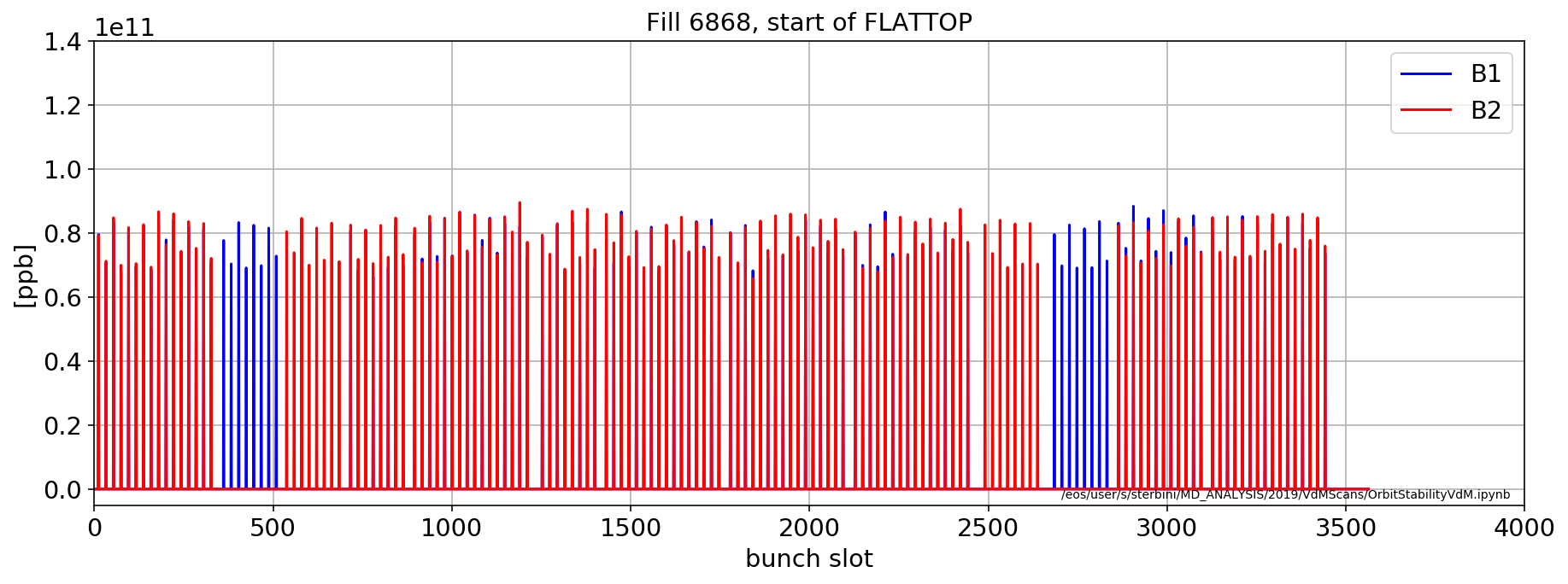
aux=importData.LHCCals2pd(['LHC.BCTFR.A6R4.B%:BUNCH_INTENSITY'],myFill, 'FLATTOP',flag='next')
plt.plot(aux['LHC.BCTFR.A6R4.B1:BUNCH_INTENSITY'].dropna().values[0],'b',label='B1')
plt.plot(aux['LHC.BCTFR.A6R4.B2:BUNCH_INTENSITY'].dropna().values[0],'r',label='B2')
plt.xlim([0,4000])
plt.title('Fill '+str(myFill)+', start of FLATTOP')
plt.xlabel('bunch slot')
plt.ylabel('[ppb]')
plt.ylim(-0.05e11, 1.4e11)
plt.grid('on')
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
plt.legend(loc='best')
<matplotlib.legend.Legend at 0x7fe80059f0f0>
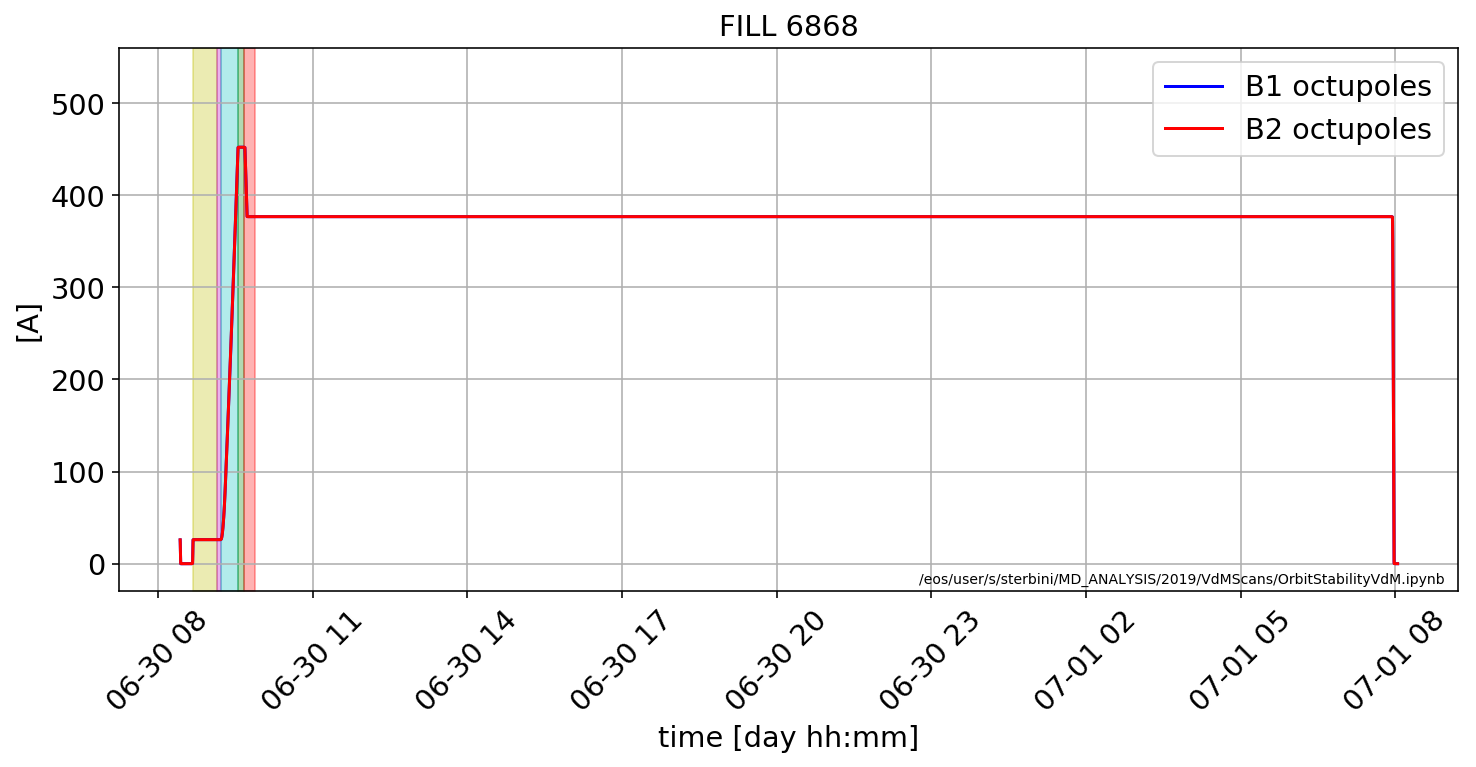
Octupoles
importData.LHCCals2pd(['RPMBB.RR17.ROF.A12B1:I_MEAS', 'RPMBB.RR17.ROF.A12B2:I_MEAS'],myFill, ['STABLE'], flag='last')
| RPMBB.RR17.ROF.A12B1:I_MEAS | RPMBB.RR17.ROF.A12B2:I_MEAS | |
|---|---|---|
| 2018-06-30 09:52:45.160000086+00:00 | 376.65 | 376.65 |
plt.figure(figsize=(12,5))
fillDF=importData.LHCFillsByNumber(myFill)
aux=importData.LHCCals2pd(['RPMBB.RR17.ROF.A12B1:I_MEAS', 'RPMBB.RR17.ROF.A12B2:I_MEAS'],myFill)
plt.plot(aux['RPMBB.RR17.ROF.A12B1:I_MEAS'].dropna(),'b',label='B1 octupoles')
plt.plot(aux['RPMBB.RR17.ROF.A12B2:I_MEAS'].dropna(),'r',label='B2 octupoles')
plt.xticks(rotation=45);
plt.xlabel('time [day hh:mm]')
plt.ylabel('[A]')
plt.title('FILL '+str(myFill))
plt.grid()
plt.legend(loc='best')
plt.ylim(-30,560)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='INJPHYS'], color='y',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='PRERAMP'], color='m',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='RAMP'], color='c',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='FLATTOP'], color='g',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='SQUEEZE'], color='orange',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='ADJUST'], color='r',alpha=.3)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
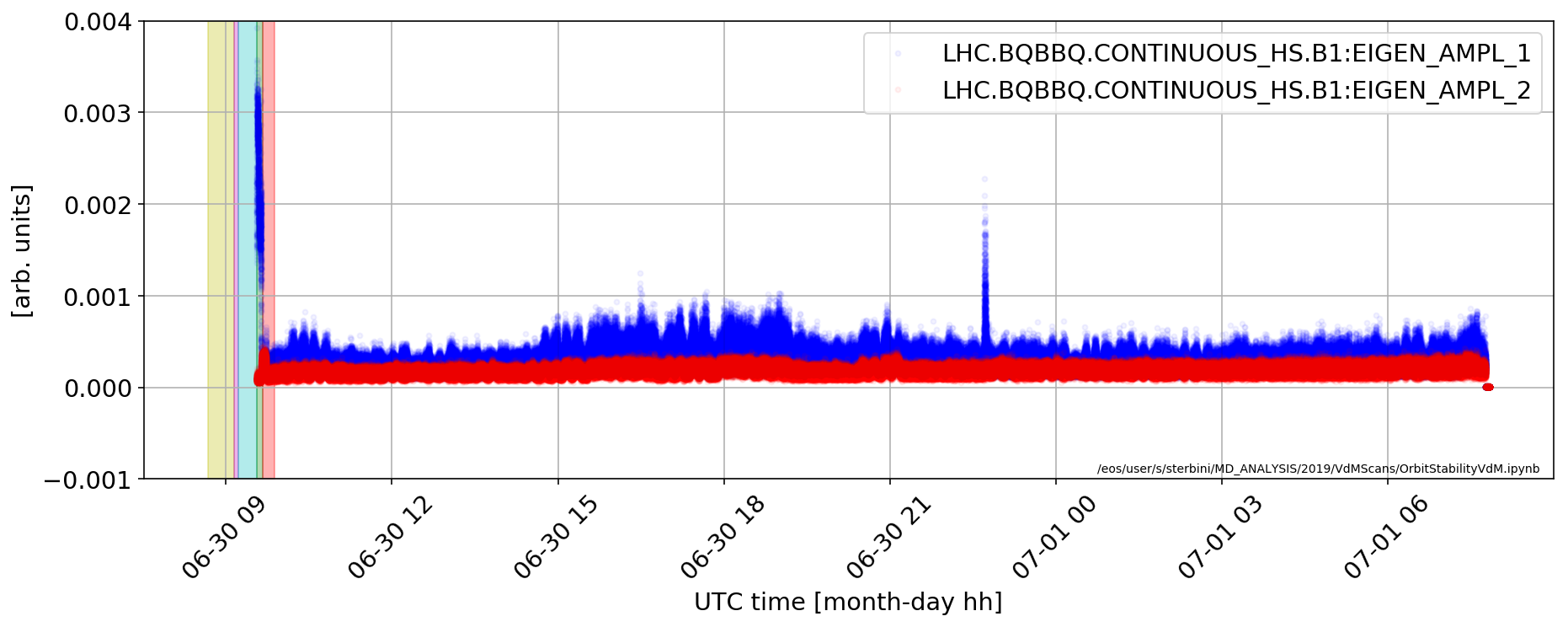
Beam stability
if 1:
myDF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BBQ.pickle')
else:
myDF=importData.LHCCals2pd(['LHC.BQBBQ.CONTINUOUS_HS.B%:EIGEN_AMPL%'],[myFill],['FLATTOP','ADJUST','STABLE'])
myDF.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BBQ.pickle')
aux=myDF#.between_time('10:08','10:27')
plt.figure(figsize=(15,5))
plt.plot(aux['LHC.BQBBQ.CONTINUOUS_HS.B1:EIGEN_AMPL_1'].dropna(),'b.',alpha=.05)
plt.plot(aux['LHC.BQBBQ.CONTINUOUS_HS.B1:EIGEN_AMPL_2'].dropna(),'r.',alpha=.05)
plt.grid(True)
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[arb. units]');
plt.legend(loc='best')
plt.ylim(-.001,.004)
plt.xticks(rotation=45)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='INJPHYS'], color='y',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='PRERAMP'], color='m',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='RAMP'], color='c',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='FLATTOP'], color='g',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='SQUEEZE'], color='orange',alpha=.3)
plotFunctions.shadedDF(plt.gca(), fillDF[fillDF['mode']=='ADJUST'], color='r',alpha=.3)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
Crossing angle configuration
importData.LHCCals2pd('LHC.RUNCONFIG:IP%-XING-%-MURAD',myFill, ['STABLE'],flag='last').describe().loc['mean']
LHC.RUNCONFIG:IP1-XING-V-MURAD 0.0
LHC.RUNCONFIG:IP2-XING-V-MURAD 145.0
LHC.RUNCONFIG:IP5-XING-H-MURAD 0.0
LHC.RUNCONFIG:IP8-XING-H-MURAD -300.0
Name: mean, dtype: float64
importData.LHCCals2pd('LHC.STATS:BETA_STAR_%',myFill, ['STABLE'],flag='last').describe().loc['mean']
LHC.STATS:BETA_STAR_ALICE 19.0
LHC.STATS:BETA_STAR_ATLAS 19.0
LHC.STATS:BETA_STAR_CMS 19.0
LHC.STATS:BETA_STAR_LHCB 24.0
Name: mean, dtype: float64
Status of the feedbacks
(importData.LHCCals2pd('LHC.BOFSU:%FB%STATE',myFill, ['RAMP'],flag='last').describe()).loc['mean']
LHC.BOFSU:CHROMA_FB_B1_STATE 0.0
LHC.BOFSU:CHROMA_FB_B2_STATE 0.0
LHC.BOFSU:COUPLING_FB_B1_STATE 0.0
LHC.BOFSU:COUPLING_FB_B2_STATE 0.0
LHC.BOFSU:ENERGY_FB_STATE 1.0
LHC.BOFSU:ORBIT_FB_STATE 1.0
LHC.BOFSU:RADIALLOOP_FB_STATE 1.0
LHC.BOFSU:TUNE_FB_B1_STATE 1.0
LHC.BOFSU:TUNE_FB_B2_STATE 1.0
Name: mean, dtype: float64
(importData.LHCCals2pd('LHC.BOFSU:%FB%STATE',myFill, ['STABLE'],flag='last').describe()).loc['mean']
LHC.BOFSU:CHROMA_FB_B1_STATE 0.0
LHC.BOFSU:CHROMA_FB_B2_STATE 0.0
LHC.BOFSU:COUPLING_FB_B1_STATE 0.0
LHC.BOFSU:COUPLING_FB_B2_STATE 0.0
LHC.BOFSU:ENERGY_FB_STATE 0.0
LHC.BOFSU:ORBIT_FB_STATE 0.0
LHC.BOFSU:RADIALLOOP_FB_STATE 0.0
LHC.BOFSU:TUNE_FB_B1_STATE 0.0
LHC.BOFSU:TUNE_FB_B2_STATE 0.0
Name: mean, dtype: float64
Orbit analysis
In LHC there are 544 BPMs x 2 planes x 2 beams (2176 BPMs in total).
The way this information is stored in CALS is optimized for data space but can be problematic in terms of post-processing.
You can retrieve the name, status and mask of all the BPMS from CALS.
The s-position of the BPM is not available in CALS and has to be derived from MAD-X.
if 1:
myBPM=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/myBPM.pickle')
else:
myBPM=importData.LHCCals2pd(['LHC.BOFSU:BPM_MASK_%', 'LHC.BOFSU:BPM_NAMES_%', 'LHC.BOFSU:BPM_STATUS_%'],myFill, ['STABLE'],flag='last')
myBPM.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/myBPM.pickle')
# There are 1088 H BPMs in total and 544 per beam
# There are 1088 V BPMs in total and 544 per beam
for i in ['LHC.BOFSU:BPM_NAMES_H', 'LHC.BOFSU:BPM_NAMES_V']:
aux=myBPM[i].dropna().iloc[0]
len([s for s in aux if 'B2' in s])
print(f'Number of {i} is {len(aux)}.')
Number of LHC.BOFSU:BPM_NAMES_H is 1088.
Number of LHC.BOFSU:BPM_NAMES_V is 1088.
# There are 1088 H BPMs in total and 544 per beam
# There are 1088 V BPMs in total and 544 per beam
for i in ['LHC.BOFSU:BPM_STATUS_H', 'LHC.BOFSU:BPM_STATUS_V']:
aux=myBPM[i].dropna().iloc[0]
print(f'Number of {i} is {len(aux)}.')
Number of LHC.BOFSU:BPM_STATUS_H is 1088.
Number of LHC.BOFSU:BPM_STATUS_V is 1088.
# There are 1088 H BPMs in total and 544 per beam
# There are 1088 V BPMs in total and 544 per beam
for i in ['LHC.BOFSU:BPM_MASK_H', 'LHC.BOFSU:BPM_MASK_V']:
aux=myBPM[i].dropna().iloc[0]
print(f'Number of {i} is {len(aux)}.')
Number of LHC.BOFSU:BPM_MASK_H is 1088.
Number of LHC.BOFSU:BPM_MASK_V is 1088.
if 1:
myBPMreadings=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/myBPMreadings.pickle')
else:
myBPMreadings=importData.LHCCals2pd('LHC.BOFSU:%POSITIONS_%',myFill, ['STABLE'],split=8)
myBPMreadings.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/myBPMreadings.pickle')
# There are 1088 H BPMs in total and 544 per beam
# There are 1088 V BPMs in total and 544 per beam
for i in ['LHC.BOFSU:POSITIONS_H', 'LHC.BOFSU:POSITIONS_V']:
aux=myBPMreadings[i].dropna().iloc[0]
print(f'Number of {i} is {len(aux)}.')
Number of LHC.BOFSU:POSITIONS_H is 1088.
Number of LHC.BOFSU:POSITIONS_V is 1088.
# Now we need to correlate the LHC.BOFSU:BPM_NAMES_H with the s-position
b1DF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/b1DF.pickle')
b2DF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/b2DF.pickle')
myS_B1_H=[]
for i in range(544):
aux=b1DF[b1DF['name'].str.contains(str.lower(myBPM['LHC.BOFSU:BPM_NAMES_H'].dropna().iloc[0][i]))]['s'].unique()
if len(aux)!=1:
print(myBPM['LHC.BOFSU:BPM_NAMES_H'].dropna().iloc[0][0])
else:
myS_B1_H.append(aux[0])
myS_B1_V=[]
for i in range(544):
aux=b1DF[b1DF['name'].str.contains(str.lower(myBPM['LHC.BOFSU:BPM_NAMES_V'].dropna().iloc[0][i]))]['s'].unique()
if len(aux)!=1:
print(myBPM['LHC.BOFSU:BPM_NAMES_V'].dropna().iloc[0][0])
else:
myS_B1_V.append(aux[0])
# the same for B2
myS_B2_H=[]
for i in range(544,1088):
aux=b2DF[b2DF['name'].str.contains(str.lower(myBPM['LHC.BOFSU:BPM_NAMES_H'].dropna().iloc[0][i]))]['s'].unique()
if len(aux)!=1:
print(myBPM['LHC.BOFSU:BPM_NAMES_H'].dropna().iloc[0][i])
else:
myS_B2_H.append(aux[0])
myS_B2_V=[]
for i in range(544,1088):
aux=b2DF[b2DF['name'].str.contains(str.lower(myBPM['LHC.BOFSU:BPM_NAMES_V'].dropna().iloc[0][i]))]['s'].unique()
if len(aux)!=1:
print(myBPM['LHC.BOFSU:BPM_NAMES_V'].dropna().iloc[0][i])
else:
myS_B2_V.append(aux[0])
# Let us consider B1 and the H/V plane
df_B1_H=pd.DataFrame([myBPM['LHC.BOFSU:BPM_NAMES_H'].dropna().iloc[0][0:544], myS_B1_H,myBPM['LHC.BOFSU:BPM_MASK_H'].dropna().iloc[0][0:544]],index=['BPM name','s [m]','mask']).transpose()
df_B1_V=pd.DataFrame([myBPM['LHC.BOFSU:BPM_NAMES_V'].dropna().iloc[0][0:544], myS_B1_V,myBPM['LHC.BOFSU:BPM_MASK_V'].dropna().iloc[0][0:544]],index=['BPM name','s [m]','mask']).transpose()
display(df_B1_H.head())
display(df_B1_V.head())
| BPM name | s [m] | mask | |
|---|---|---|---|
| 0 | BPMSW.1R1.B1 | 21.564 | 1 |
| 1 | BPMWF.A1R1.B1 | 21.724 | 0 |
| 2 | BPMS.2R1.B1 | 31.529 | 1 |
| 3 | BPMSY.4R1.B1 | 58.3145 | 1 |
| 4 | BPMWB.4R1.B1 | 151.094 | 1 |
| BPM name | s [m] | mask | |
|---|---|---|---|
| 0 | BPMSW.1R1.B1 | 21.564 | 1 |
| 1 | BPMWF.A1R1.B1 | 21.724 | 0 |
| 2 | BPMS.2R1.B1 | 31.529 | 1 |
| 3 | BPMSY.4R1.B1 | 58.3145 | 1 |
| 4 | BPMWB.4R1.B1 | 151.094 | 1 |
# Let us consider B2 and the H/V plane
df_B2_H=pd.DataFrame([myBPM['LHC.BOFSU:BPM_NAMES_H'].dropna().iloc[0][544:1088], myS_B2_H , myBPM['LHC.BOFSU:BPM_MASK_H'].dropna().iloc[0][544:1088]],index=['BPM name','s [m]','mask']).transpose()
df_B2_V=pd.DataFrame([myBPM['LHC.BOFSU:BPM_NAMES_V'].dropna().iloc[0][544:1088], myS_B2_V, myBPM['LHC.BOFSU:BPM_MASK_V'].dropna().iloc[0][544:1088]],index=['BPM name','s [m]','mask']).transpose()
display(df_B2_H.head())
display(df_B2_V.head())
| BPM name | s [m] | mask | |
|---|---|---|---|
| 0 | BPMSW.1R1.B2 | 21.564 | 1 |
| 1 | BPMWF.A1R1.B2 | 21.724 | 0 |
| 2 | BPMS.2R1.B2 | 31.529 | 1 |
| 3 | BPMSY.4R1.B2 | 58.3145 | 1 |
| 4 | BPMWB.4R1.B2 | 151.17 | 1 |
| BPM name | s [m] | mask | |
|---|---|---|---|
| 0 | BPMSW.1R1.B2 | 21.564 | 1 |
| 1 | BPMWF.A1R1.B2 | 21.724 | 0 |
| 2 | BPMS.2R1.B2 | 31.529 | 1 |
| 3 | BPMSY.4R1.B2 | 58.3145 | 1 |
| 4 | BPMWB.4R1.B2 | 151.17 | 1 |
# IMPORTANT: from array to pandas columns
# B1H
myBPM_B1_H=pd.DataFrame(myBPMreadings['LHC.BOFSU:POSITIONS_H'].copy())
myBPM_B1_H.head()
aux=df_B1_H[df_B1_H['mask']==1]['BPM name']
for i,j in zip(aux.values, aux.index):
myBPM_B1_H[i]=myBPM_B1_H['LHC.BOFSU:POSITIONS_H'].apply(lambda x: x[j])
del myBPM_B1_H['LHC.BOFSU:POSITIONS_H']
# IMPORTANT: from array to pandas columns
# B1V
myBPM_B1_V=pd.DataFrame(myBPMreadings['LHC.BOFSU:POSITIONS_V'].copy())
myBPM_B1_V.head()
aux=df_B1_H[df_B1_H['mask']==1]['BPM name']
for i,j in zip(aux.values, aux.index):
myBPM_B1_V[i]=myBPM_B1_V['LHC.BOFSU:POSITIONS_V'].apply(lambda x: x[j])
del myBPM_B1_V['LHC.BOFSU:POSITIONS_V']
# IMPORTANT: from array to pandas columns
# B2H
myBPM_B2_H=pd.DataFrame(myBPMreadings['LHC.BOFSU:POSITIONS_H'].copy())
myBPM_B2_H.head()
aux=df_B2_H[df_B2_H['mask']==1]['BPM name']
for i,j in zip(aux.values, aux.index):
myBPM_B2_H[i]=myBPM_B2_H['LHC.BOFSU:POSITIONS_H'].apply(lambda x: x[j+544])
del myBPM_B2_H['LHC.BOFSU:POSITIONS_H']
# IMPORTANT: from array to pandas columns
# B2V
myBPM_B2_V=pd.DataFrame(myBPMreadings['LHC.BOFSU:POSITIONS_V'].copy())
myBPM_B2_V.head()
aux=df_B2_V[df_B2_V['mask']==1]['BPM name']
for i,j in zip(aux.values, aux.index):
myBPM_B2_V[i]=myBPM_B2_V['LHC.BOFSU:POSITIONS_V'].apply(lambda x: x[j+544])
del myBPM_B2_V['LHC.BOFSU:POSITIONS_V']
Important results
We have now four tables for the time vatiation of the H/V plane in B½.
for i in [myBPM_B1_H,myBPM_B1_V,myBPM_B2_H,myBPM_B2_V]:
display(i.head(3))
| BPMSW.1R1.B1 | BPMS.2R1.B1 | BPMSY.4R1.B1 | BPMWB.4R1.B1 | BPMYA.4R1.B1 | BPM.5R1.B1 | BPMR.6R1.B1 | BPMSX.7R1.B1 | BPM.8R1.B1 | BPM.9R1.B1 | ... | BPM.9L1.B1 | BPM.8L1.B1 | BPMR.7L1.B1 | BPM.6L1.B1 | BPMR.5L1.B1 | BPMYA.4L1.B1 | BPMWB.4L1.B1 | BPMSY.4L1.B1 | BPMS.2L1.B1 | BPMSW.1L1.B1 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2018-06-30 09:52:46.567000151+00:00 | -506.99 | -191.17 | -379.43 | 426.54 | 137.41 | 138.58 | 16.29 | -131.38 | 438.92 | -72.87 | ... | 269.33 | -377.80 | 490.24 | 315.39 | 222.96 | -67.14 | 297.23 | -1053.89 | 66.77 | 22.62 |
| 2018-06-30 09:52:47.526999950+00:00 | -507.76 | -190.33 | -380.41 | 425.07 | 136.62 | 137.36 | 14.53 | -130.32 | 439.50 | -74.21 | ... | 268.83 | -377.71 | 489.68 | 314.90 | 223.15 | -67.16 | 296.13 | -1054.74 | 67.28 | 20.67 |
| 2018-06-30 09:52:48.526999950+00:00 | -506.38 | -192.56 | -379.26 | 426.79 | 136.76 | 138.20 | 15.56 | -129.11 | 439.75 | -75.11 | ... | 268.82 | -380.25 | 490.24 | 314.18 | 223.29 | -67.59 | 298.66 | -1054.54 | 68.91 | 21.14 |
3 rows × 518 columns
| BPMSW.1R1.B1 | BPMS.2R1.B1 | BPMSY.4R1.B1 | BPMWB.4R1.B1 | BPMYA.4R1.B1 | BPM.5R1.B1 | BPMR.6R1.B1 | BPMSX.7R1.B1 | BPM.8R1.B1 | BPM.9R1.B1 | ... | BPM.9L1.B1 | BPM.8L1.B1 | BPMR.7L1.B1 | BPM.6L1.B1 | BPMR.5L1.B1 | BPMYA.4L1.B1 | BPMWB.4L1.B1 | BPMSY.4L1.B1 | BPMS.2L1.B1 | BPMSW.1L1.B1 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2018-06-30 09:52:46.567000151+00:00 | -149.91 | -160.70 | -306.87 | -74.58 | 752.21 | -901.88 | -104.03 | 911.58 | 242.89 | -478.25 | ... | -228.21 | 251.39 | -375.76 | 600.20 | -569.59 | 711.48 | 152.80 | 438.11 | -208.22 | -257.47 |
| 2018-06-30 09:52:47.526999950+00:00 | -150.36 | -161.56 | -309.88 | -75.68 | 749.99 | -905.23 | -102.46 | 908.94 | 243.82 | -477.88 | ... | -227.20 | 251.49 | -374.02 | 601.05 | -568.82 | 712.39 | 155.27 | 438.42 | -207.95 | -258.24 |
| 2018-06-30 09:52:48.526999950+00:00 | -149.17 | -162.95 | -308.38 | -76.76 | 749.51 | -903.49 | -103.59 | 909.42 | 242.95 | -476.18 | ... | -227.08 | 252.49 | -372.03 | 602.15 | -568.59 | 715.00 | 158.12 | 438.50 | -204.34 | -257.92 |
3 rows × 518 columns
| BPMSW.1R1.B2 | BPMS.2R1.B2 | BPMSY.4R1.B2 | BPMWB.4R1.B2 | BPMYA.4R1.B2 | BPMR.5R1.B2 | BPM.6R1.B2 | BPMRA.7R1.B2 | BPM.8R1.B2 | BPM.9R1.B2 | ... | BPM.10L1.B2 | BPM.9L1.B2 | BPM.8L1.B2 | BPM.7L1.B2 | BPMR.6L1.B2 | BPM.5L1.B2 | BPMWB.4L1.B2 | BPMSY.4L1.B2 | BPMS.2L1.B2 | BPMSW.1L1.B2 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2018-06-30 09:52:46.567000151+00:00 | -241.29 | 38.00 | 62.46 | -1095.68 | 155.71 | 163.05 | 14.85 | 66.14 | -30.09 | -12.40 | ... | -618.24 | 132.72 | 16.05 | 569.03 | 182.15 | -27.24 | 124.67 | -1626.10 | -37.14 | 9.27 |
| 2018-06-30 09:52:47.526999950+00:00 | -241.70 | 38.37 | 64.46 | -1095.65 | 156.91 | 162.70 | 15.61 | 64.58 | -31.28 | -10.85 | ... | -619.22 | 132.62 | 16.98 | 569.32 | 183.79 | -28.06 | 125.15 | -1625.55 | -36.74 | 8.89 |
| 2018-06-30 09:52:48.526999950+00:00 | -242.74 | 36.43 | 63.62 | -1094.91 | 157.67 | 161.81 | 14.85 | 65.68 | -31.44 | -11.90 | ... | -617.91 | 131.87 | 16.73 | 569.41 | 184.00 | -27.66 | 123.23 | -1624.02 | -37.89 | 7.87 |
3 rows × 523 columns
| BPMSW.1R1.B2 | BPMS.2R1.B2 | BPMSY.4R1.B2 | BPMWB.4R1.B2 | BPMYA.4R1.B2 | BPMR.5R1.B2 | BPM.6R1.B2 | BPMRA.7R1.B2 | BPM.8R1.B2 | BPM.9R1.B2 | ... | BPM.8L1.B2 | BPM.7L1.B2 | BPMSX.7L1.B2 | BPMR.6L1.B2 | BPM.5L1.B2 | BPMYA.4L1.B2 | BPMWB.4L1.B2 | BPMSY.4L1.B2 | BPMS.2L1.B2 | BPMSW.1L1.B2 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2018-06-30 09:52:46.567000151+00:00 | 257.17 | 190.90 | -139.11 | -235.79 | 133.49 | -155.30 | -201.60 | 66.57 | 438.31 | -157.35 | ... | 248.56 | 103.21 | 95.01 | 297.41 | -291.00 | -166.64 | -377.31 | -406.43 | 194.84 | 282.24 |
| 2018-06-30 09:52:47.526999950+00:00 | 257.85 | 191.66 | -138.02 | -235.30 | 135.28 | -157.55 | -202.23 | 67.39 | 439.00 | -159.81 | ... | 249.39 | 102.72 | 97.34 | 297.08 | -294.39 | -164.71 | -376.83 | -407.38 | 195.38 | 283.36 |
| 2018-06-30 09:52:48.526999950+00:00 | 257.74 | 193.08 | -134.43 | -235.48 | 134.45 | -156.23 | -203.00 | 67.01 | 437.74 | -159.99 | ... | 249.02 | 102.78 | 94.92 | 298.63 | -294.53 | -164.25 | -379.24 | -407.70 | 194.14 | 281.54 |
3 rows × 528 columns
Plotting the machine Closed Orbit
We average it in time.
plt.plot(df_B1_H[df_B1_H['mask']==1]['s [m]'].values, myBPM_B1_H.mean()/1000,'b',label='x-plane')
plt.plot(df_B1_H[df_B1_H['mask']==1]['s [m]'].values, myBPM_B1_V.mean()/1000,'r',label='y-plane')
plt.xlabel('s [m]')
plt.ylabel('[mm]')
plt.grid(True)
plt.legend(loc='best')
<matplotlib.legend.Legend at 0x7fea87a68630>
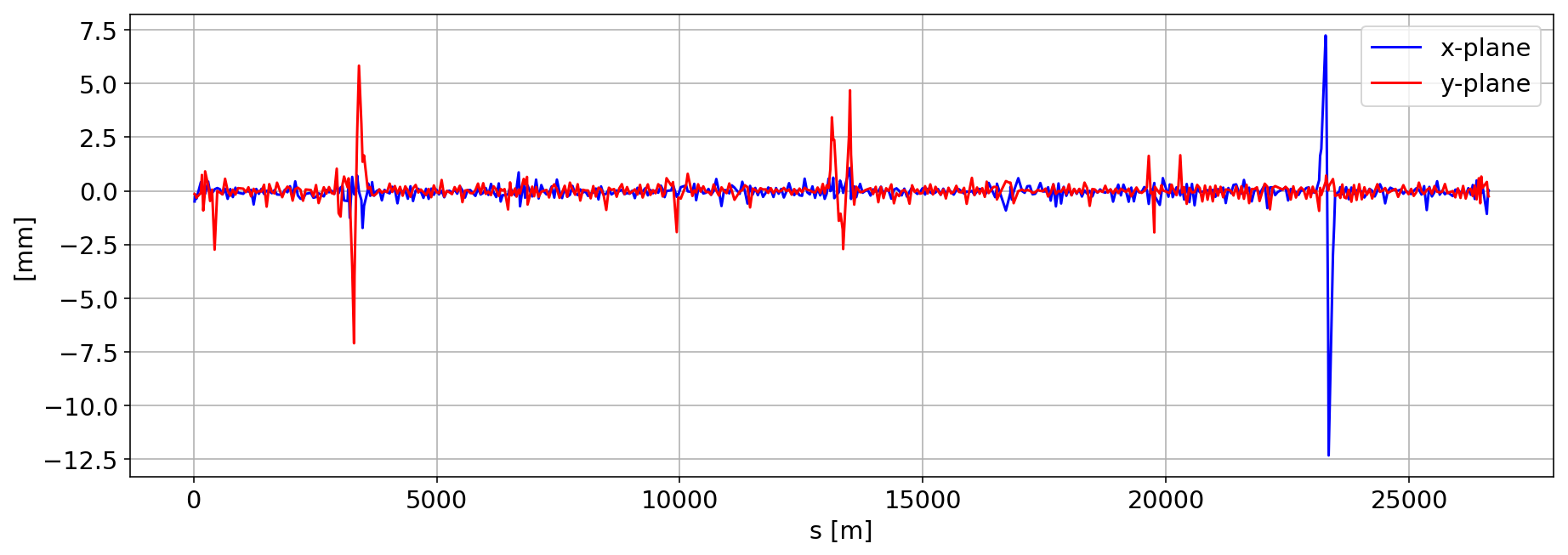
Let us focus during the IP1 H-scan
smallDF_B1=myBPM_B1_H.iloc[900:2200].copy()
plt.plot(smallDF_B1['BPMSW.1R1.B1']/1000,'b',lw=3)
smallDF_B2=myBPM_B2_H.iloc[900:2200].copy()
plt.plot(smallDF_B2['BPMSW.1R1.B2']/1000,'r',lw=3)
plt.grid(True)
plt.xlabel('time [month hh:mm]')
plt.ylabel('[mm]')
plt.grid(True)
plt.legend(loc='best')
<matplotlib.legend.Legend at 0x7fe8b2d80048>
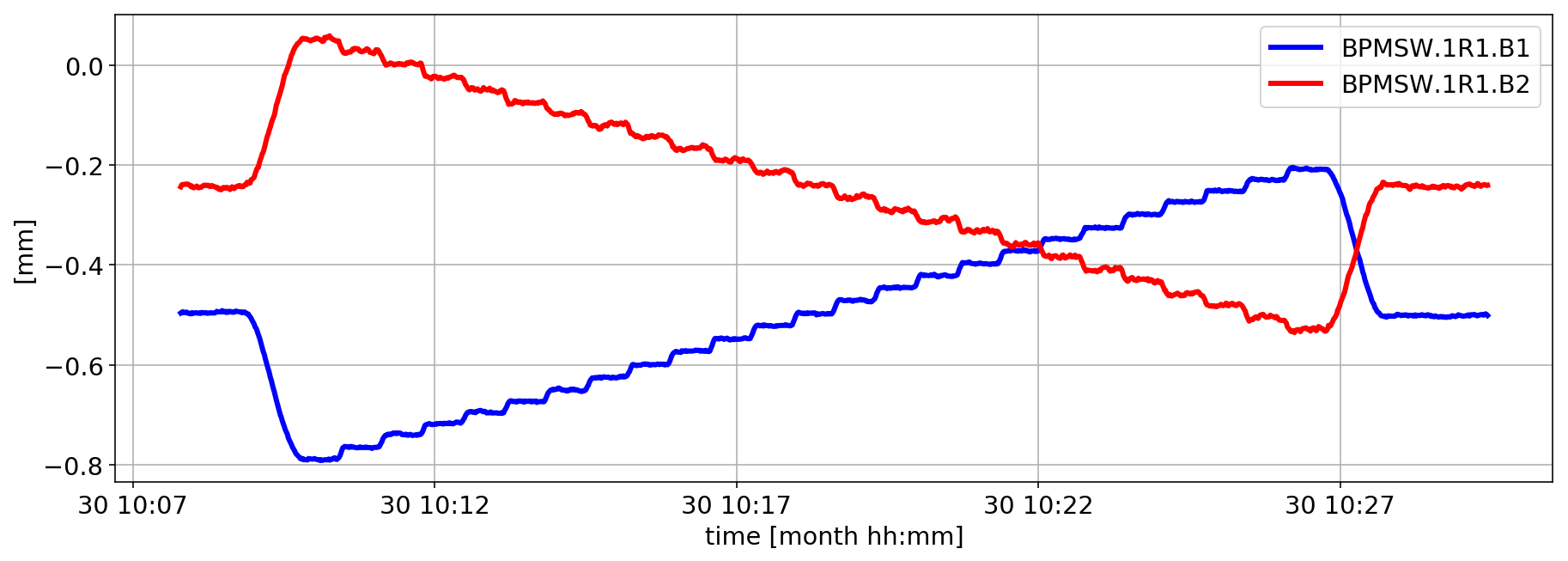
Stability consideration during the scan
if 1:
myDF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BBQ.pickle')
else:
myDF=importData.LHCCals2pd(['LHC.BQBBQ.CONTINUOUS_HS.B%:EIGEN_AMPL%'],[myFill],['FLATTOP','ADJUST','STABLE'])
myDF.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/BBQ.pickle')
aux=myDF.between_time('10:07:47','10:29:26')
plt.figure(figsize=(15,10))
plt.plot(aux['LHC.BQBBQ.CONTINUOUS_HS.B1:EIGEN_AMPL_1'].dropna(),'b.-',alpha=.5)
plt.plot(aux['LHC.BQBBQ.CONTINUOUS_HS.B1:EIGEN_AMPL_2'].dropna(),'r.-',alpha=.5)
plt.grid(True)
plt.xlabel('UTC time [month-day hh]');
plt.ylabel('[arb. units]');
plt.legend(loc='best')
ax2 = plt.gca().twinx() # instantiate a second axes that shares the same x-axis
color = 'm'
ax2.set_ylabel('BPMSW.1R1.B1 [mm]', color=color) # we already handled the x-label with ax1
ax2.plot(smallDF_B1['BPMSW.1R1.B1']/1000,'m',lw=3)
ax2.tick_params(axis='y', labelcolor=color)
#plt.ylim(-.001,.004)
plotFunctions.setSourcePlot(plt.gca(), mySource,color='k')
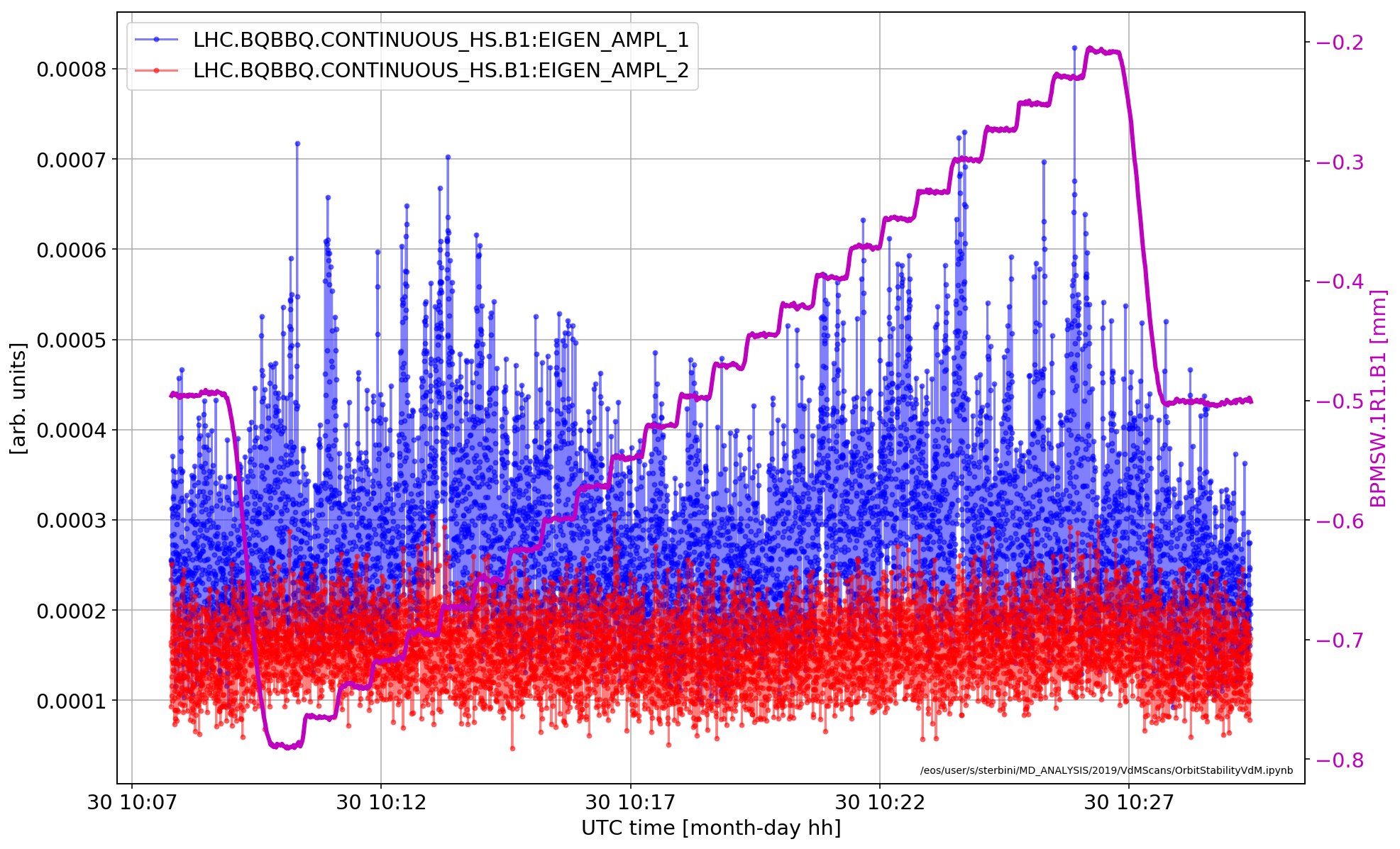
It is interesting to observe that the activity in B1 is increasing, as expected, when the two beams are not HO.
Singular value decomposition
Given the orbit matrix (time \times s-position) we perform a SVD to establish a hierarchy of the modes in the CO evolution. This is a standard procedure and is only one application of the SVD algorithm. To have an interesting overview of SVD potential, I suggest this collection of examples here.
The SVD is a generatization of the diagonal decomposition. The latter can consider only for square matrices but it is not always possible. The SVD can be done for square and rectangular matrices and is always possible.
# Example
myMatrix=np.array([[1,1],[0,1]])
U,S,V_star=np.linalg.svd(myMatrix)
print('===============')
print(f"myMatrix size is {np.shape(myMatrix)}")
display(myMatrix)
print('===============')
print(f"U size is {np.shape(U)}")
display(U)
print('===============')
print(f"S size is {np.shape(S)}")
display(S)
print('===============')
print(f"V_star size is {np.shape(V_star)}")
display(V_star)
print('===============')
print(f"Sanity check: {np.allclose(myMatrix,U * S @ V_star)}")
===============
myMatrix size is (2, 2)
array([[1, 1],
[0, 1]])
===============
U size is (2, 2)
array([[ 0.85065081, -0.52573111],
[ 0.52573111, 0.85065081]])
===============
S size is (2,)
array([1.61803399, 0.61803399])
===============
V_star size is (2, 2)
array([[ 0.52573111, 0.85065081],
[-0.85065081, 0.52573111]])
===============
Sanity check: True
D,A=np.linalg.eig(myMatrix)
print('===============')
print(f"D size is {np.shape(D)}")
display(D)
print('===============')
print(f"A size is {np.shape(A)}")
display(A)
print('===============')
print(f"Sanity check: {np.allclose(myMatrix,A @ np.diag(D) @ np.linalg.inv(A))}")
===============
D size is (2,)
array([1., 1.])
===============
A size is (2, 2)
array([[ 1.00000000e+00, -1.00000000e+00],
[ 0.00000000e+00, 2.22044605e-16]])
===============
Sanity check: False
The algorithm to compute the SVD of M is based on the diagonal form of MM^{*} and M^{*}M. The SV are the square root of the MM^{*} (M^{*}M or since they are identical). The SVD modes are related to the eigenvecotors of MM^{*} and M^{*}M.
# just to check
np.linalg.eig(myMatrix.transpose()@myMatrix)
(array([0.38196601, 2.61803399]), array([[-0.85065081, -0.52573111],
[ 0.52573111, -0.85065081]]))
# just to check
np.linalg.eig(myMatrix@myMatrix.transpose())
(array([2.61803399, 0.38196601]), array([[ 0.85065081, -0.52573111],
[ 0.52573111, 0.85065081]]))
A grephical representation of the SVD can be found on https://en.wikipedia.org

The diagonal form associated with a linear endomorphism defines a basis that is algebrically transformed, The SVD defines an orho-normal basis V that is algebrically transformed in the orho-normal basis U.
Since the SV are ordered and since the matrices U and V are orthogonal (i.e., bases of orthonormal vectors), a metric hierarchy is established.
If the matrix M represents the evolution of the orbits (s-direction) in time (t-direction), we can decompose the orbit of the machine in s-dipendent modes (V_star rows) that vary in time (U columns). The modes will be order by their metric.
# we remove the mean orbit
del U, S, V, aux
aux=smallDF_B1-smallDF_B1.mean()
# we decompose the matrix
U,S,V=np.linalg.svd(aux.values,full_matrices=False)
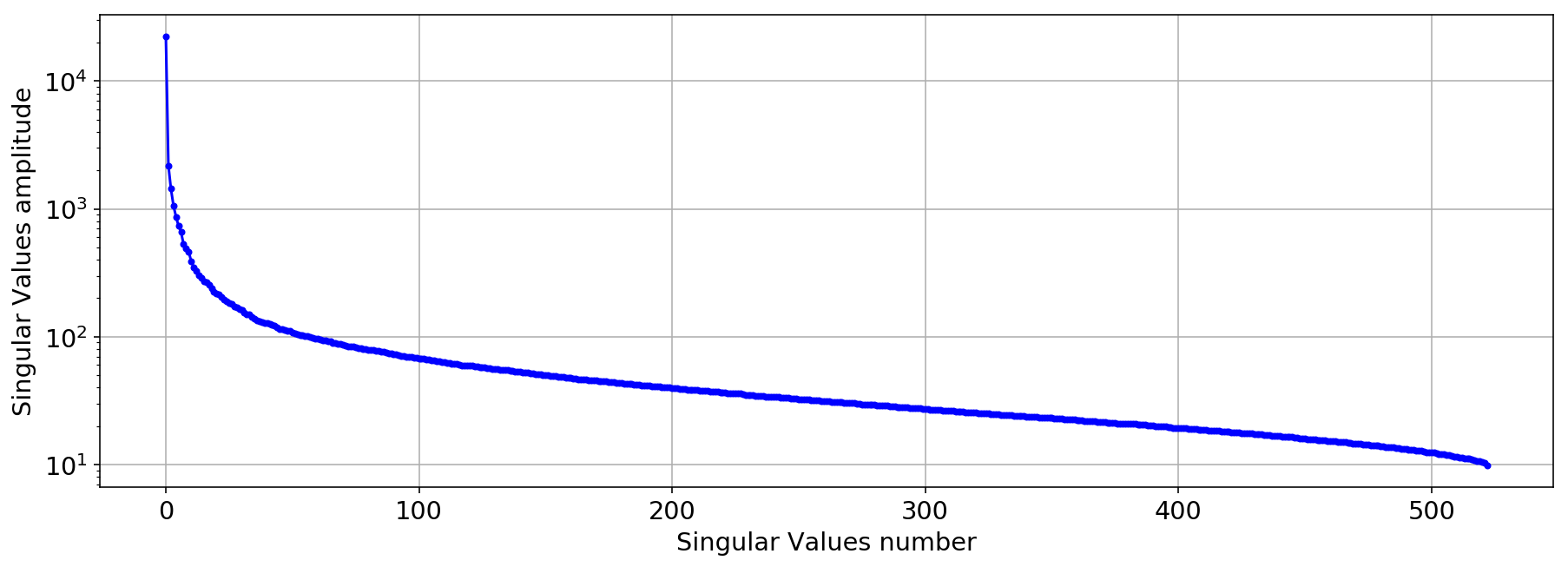
plt.semilogy(S,'.-b')
#plt.xlim([0,40])
plt.xlabel('Singular Values number')
plt.ylabel('Singular Values amplitude');
plt.grid(True)
# sanity check
print(f'Sanity check: {np.allclose(aux.values,U * S @ V, equal_nan=True)}')
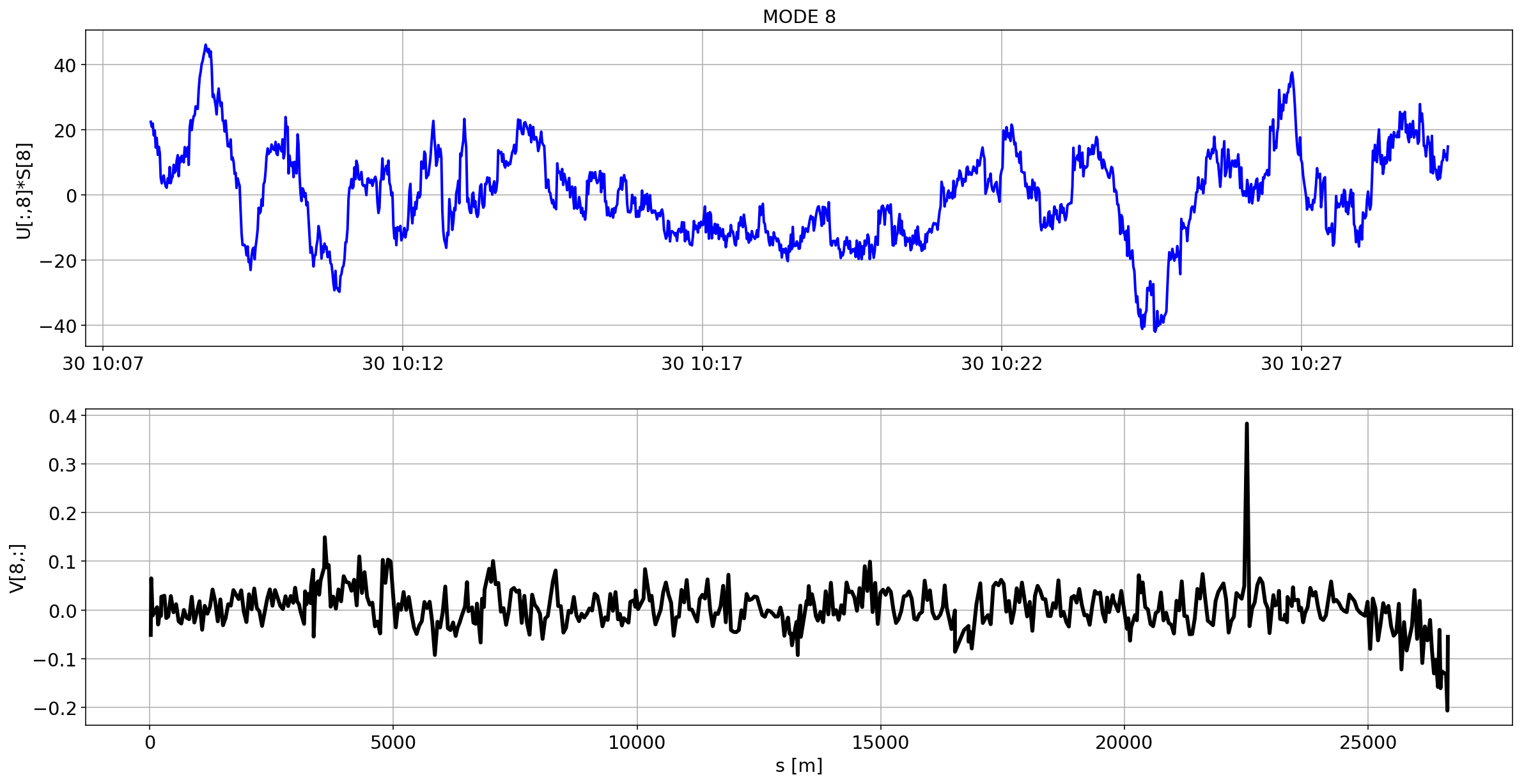
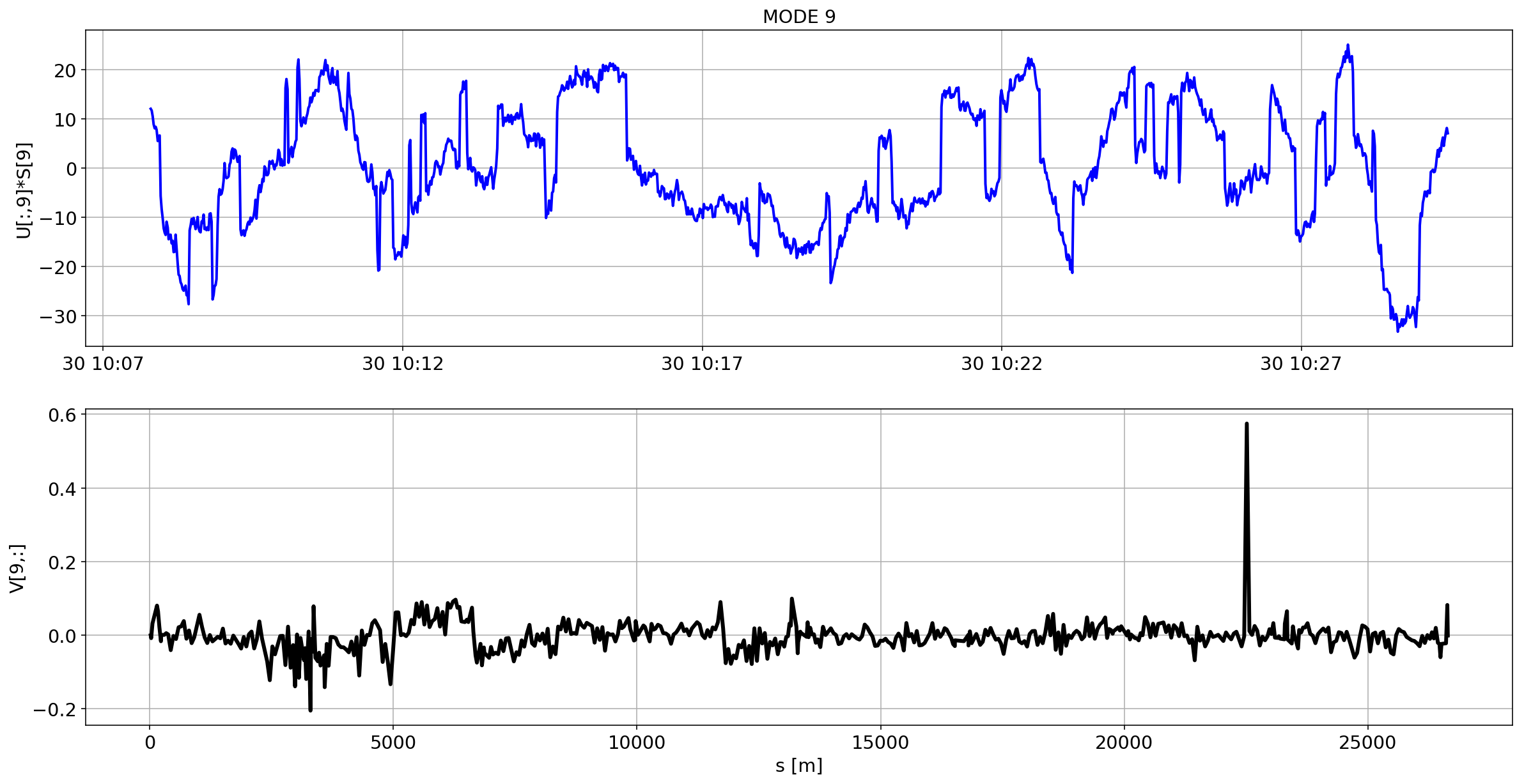
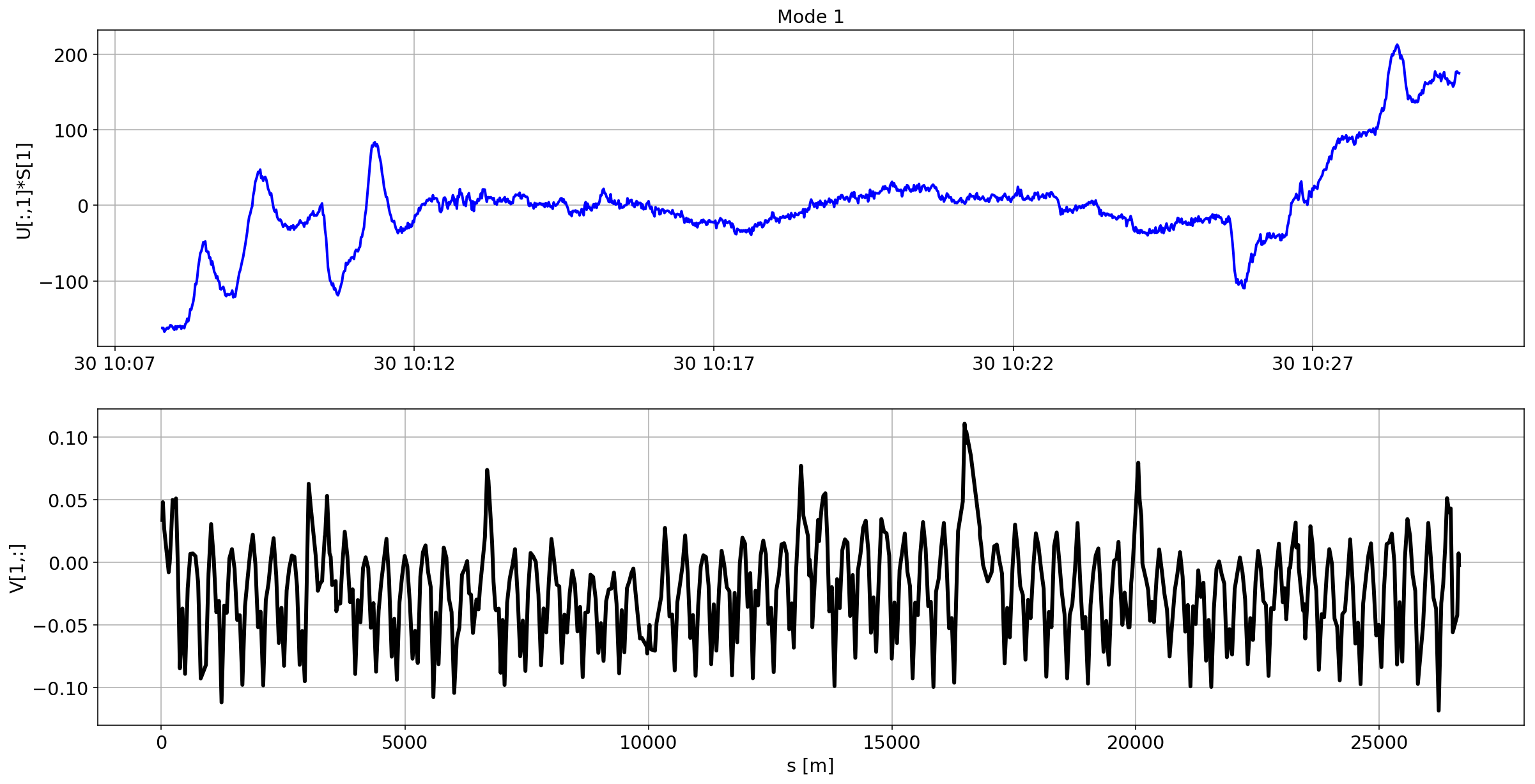
def plotMode_B1(mode=0):
plt.figure(figsize=(20,10))
plt.subplot(2,1,1)
plt.plot(aux.index,U[:,mode]*S[mode],'b',lw=2)
plt.grid(True)
plt.ylabel(f'U[:,{mode}]*S[{mode}]')
plt.title(f'MODE {mode}')
plt.subplot(2,1,2)
plt.plot(df_B1_H[df_B1_H['mask']==1]['s [m]'].values,V[mode,:],'k',lw=3)
plt.grid(True)
plt.ylabel(f'V[{mode},:]')
plt.xlabel('s [m]')
Sanity check: False
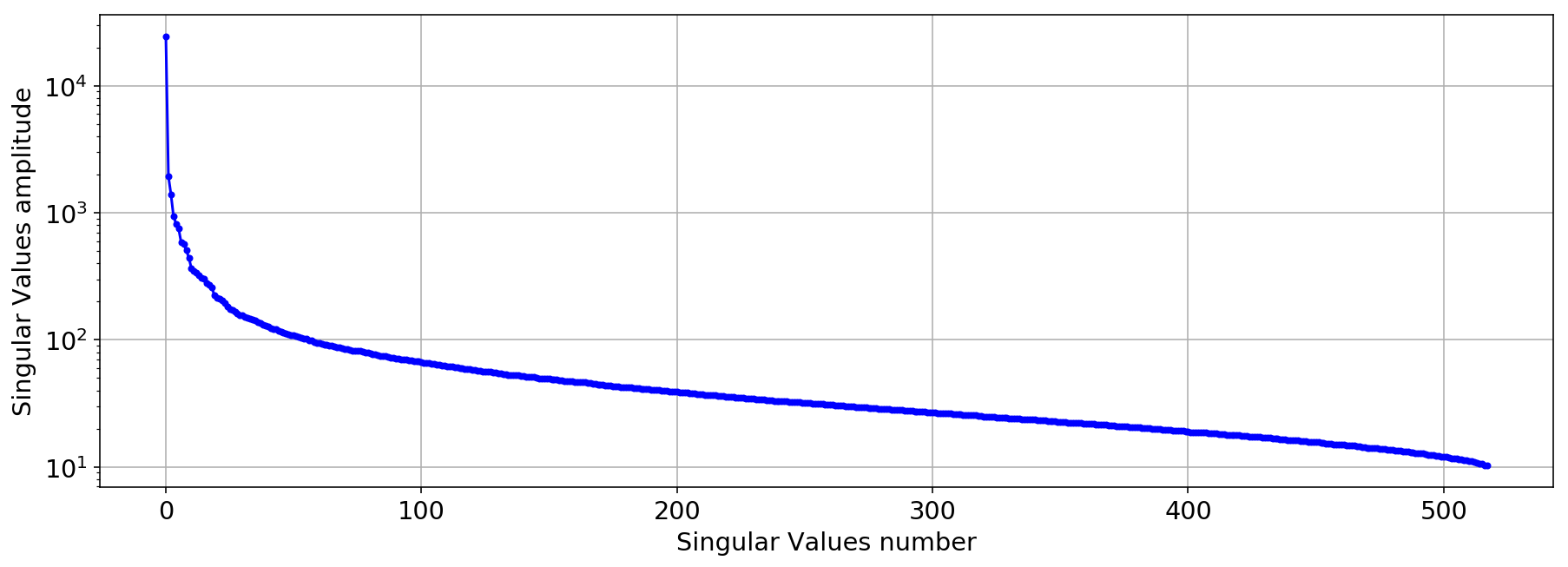
for i in range(10):
plotMode_B1(mode=i)
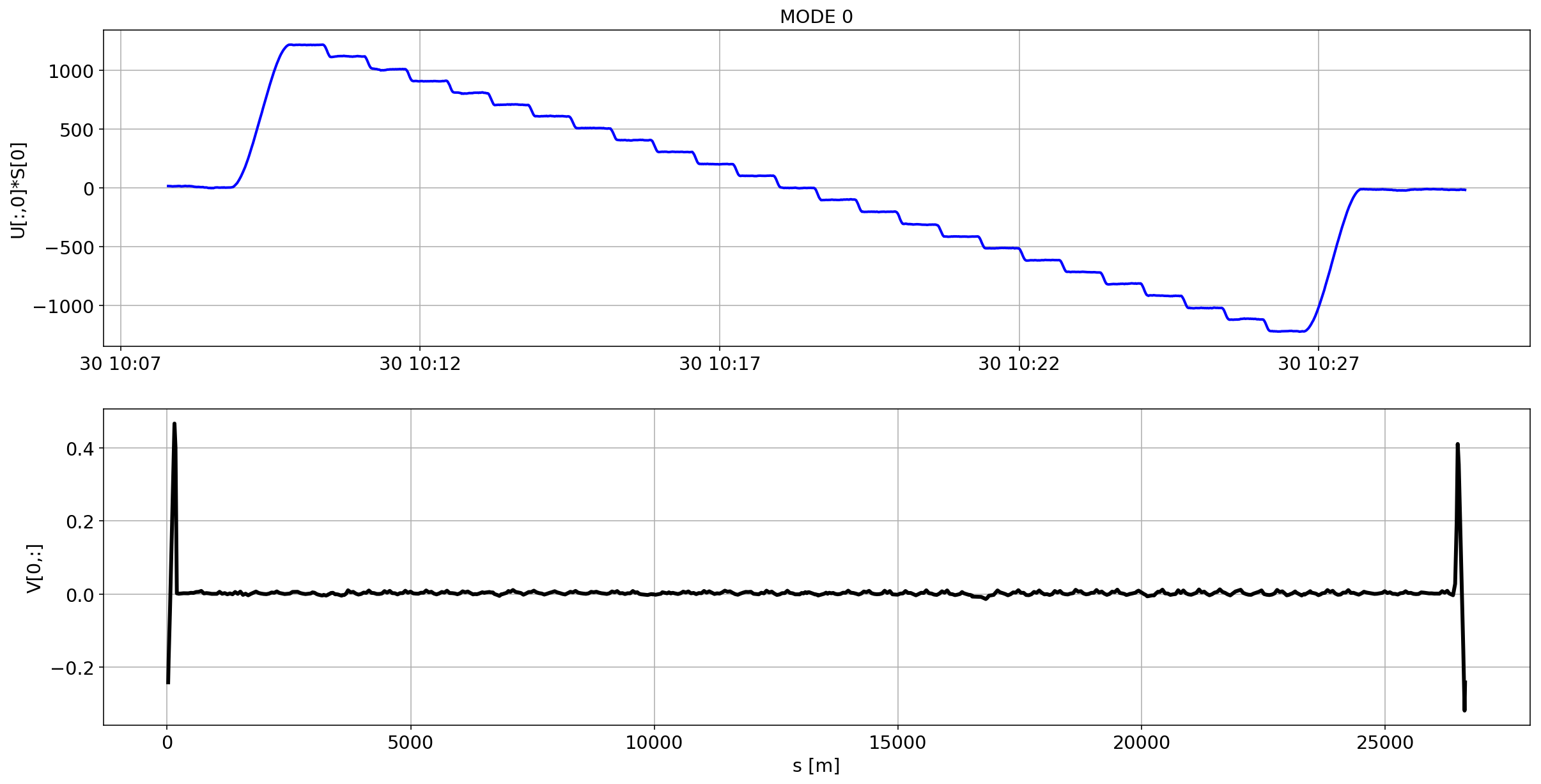
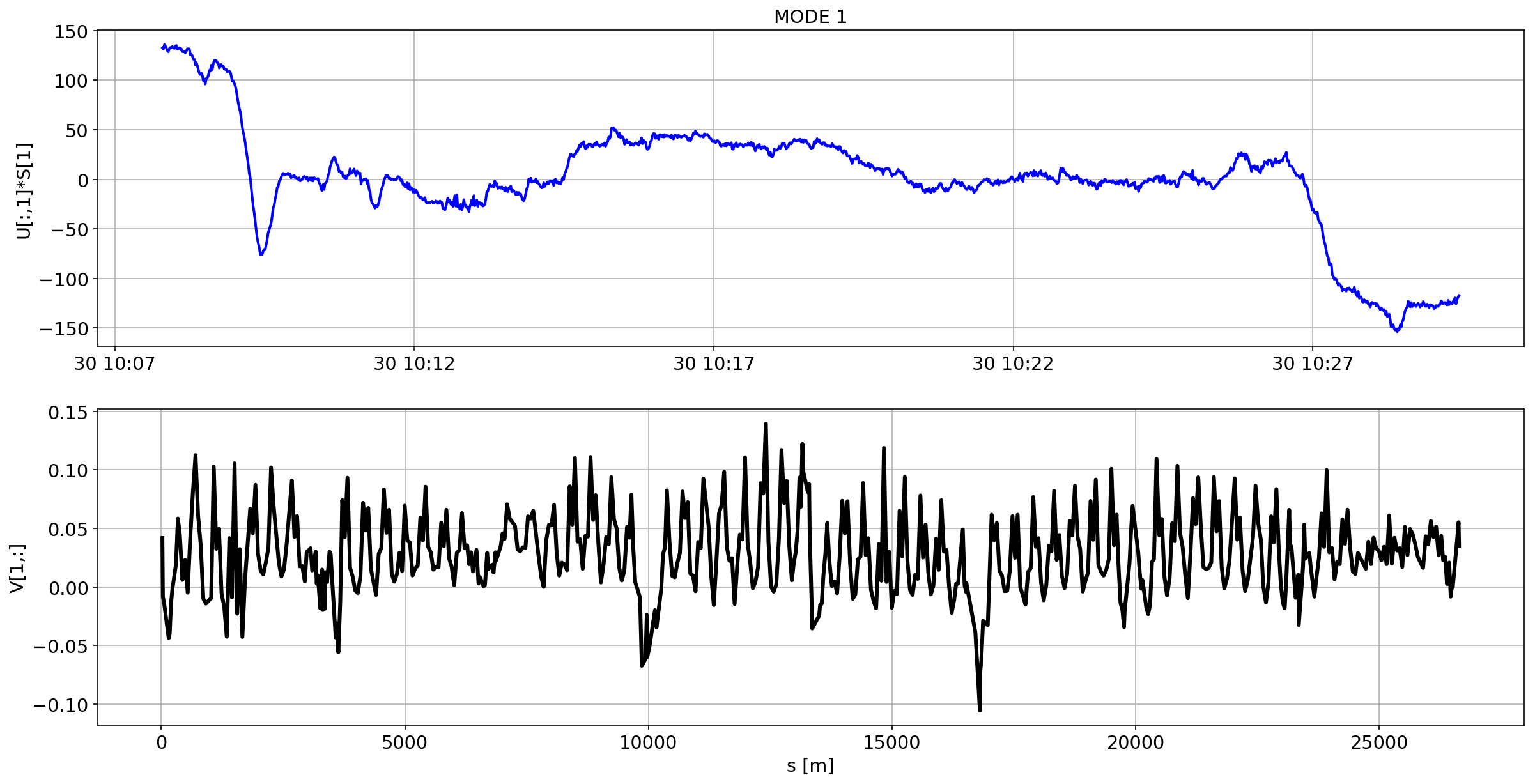
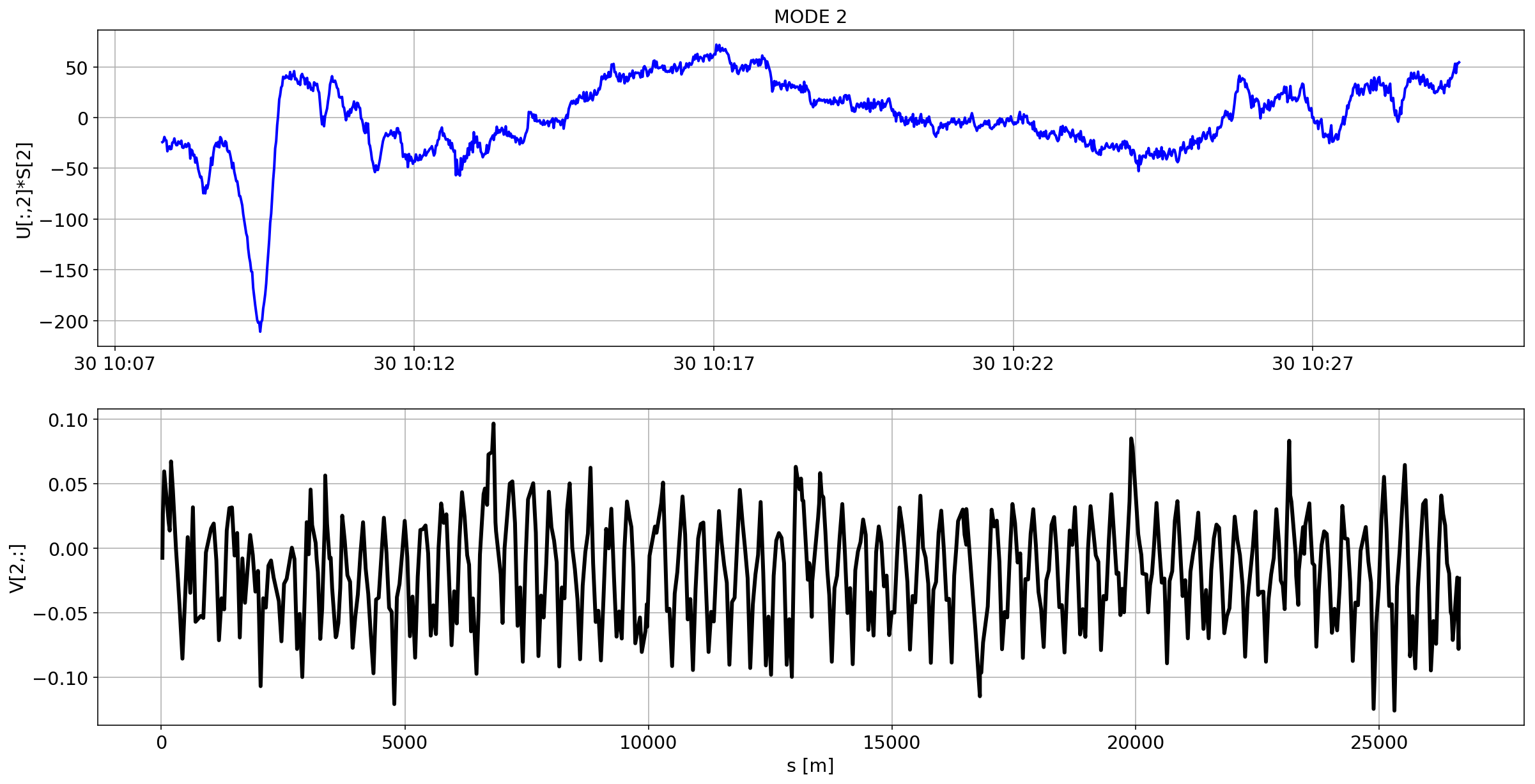
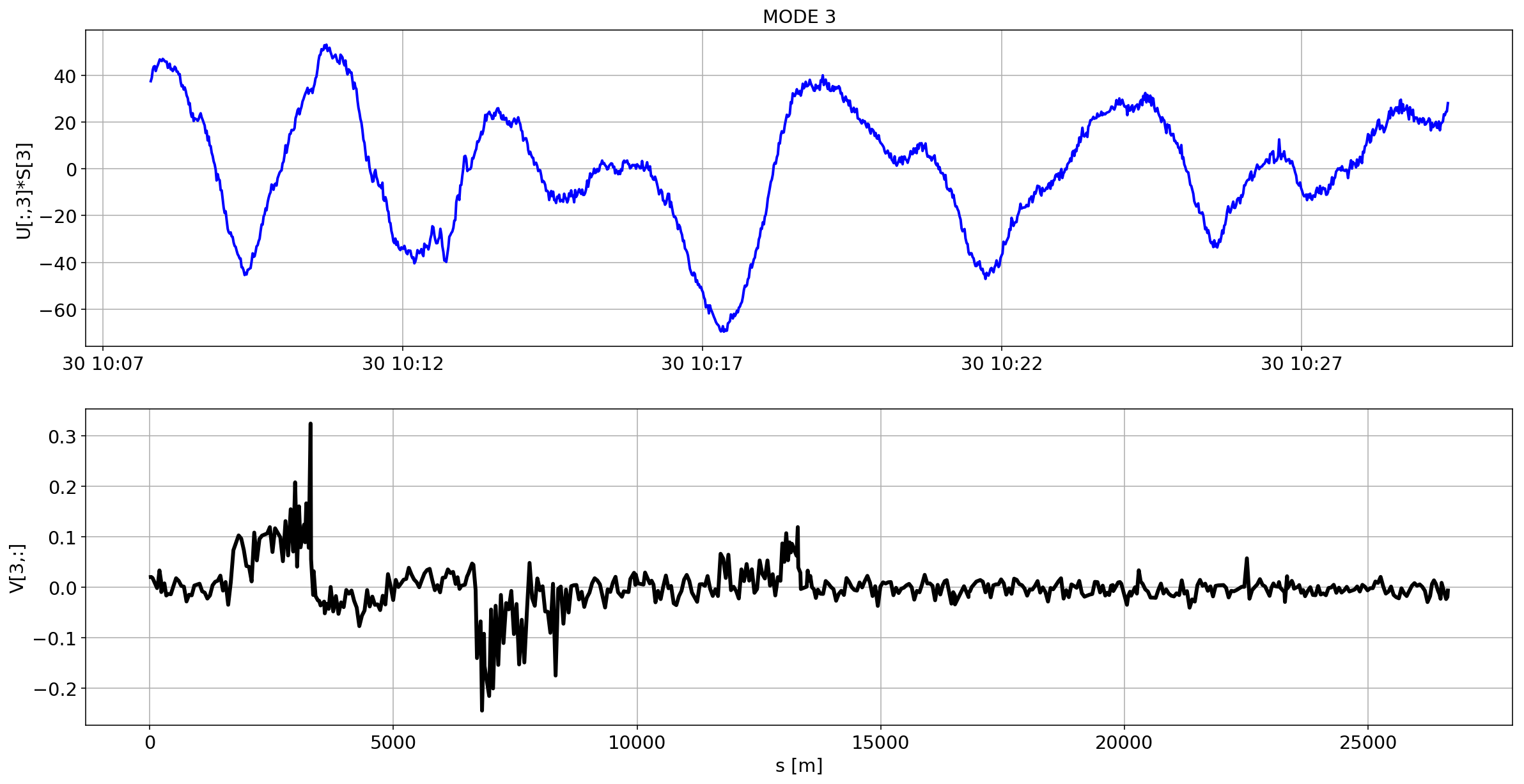
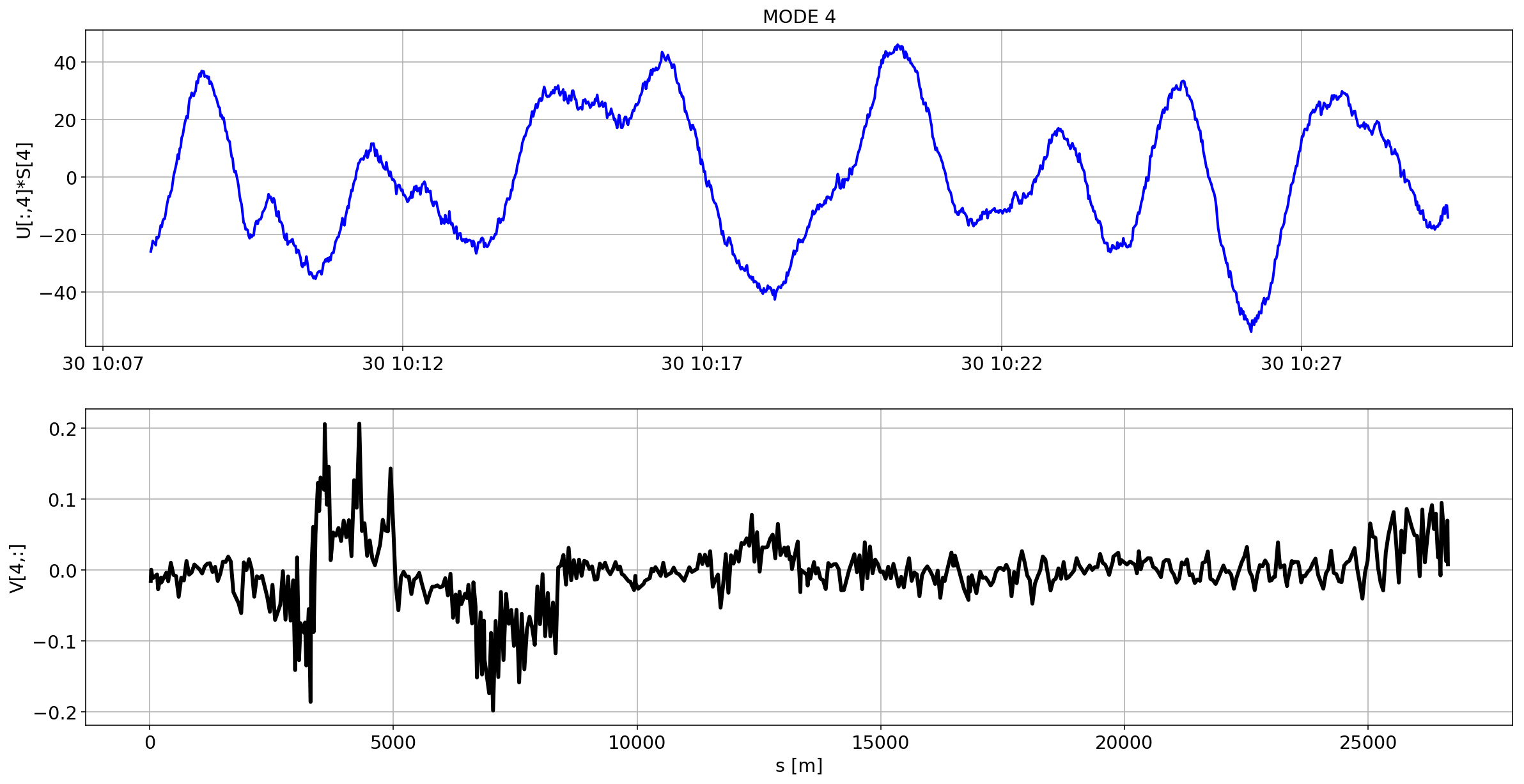
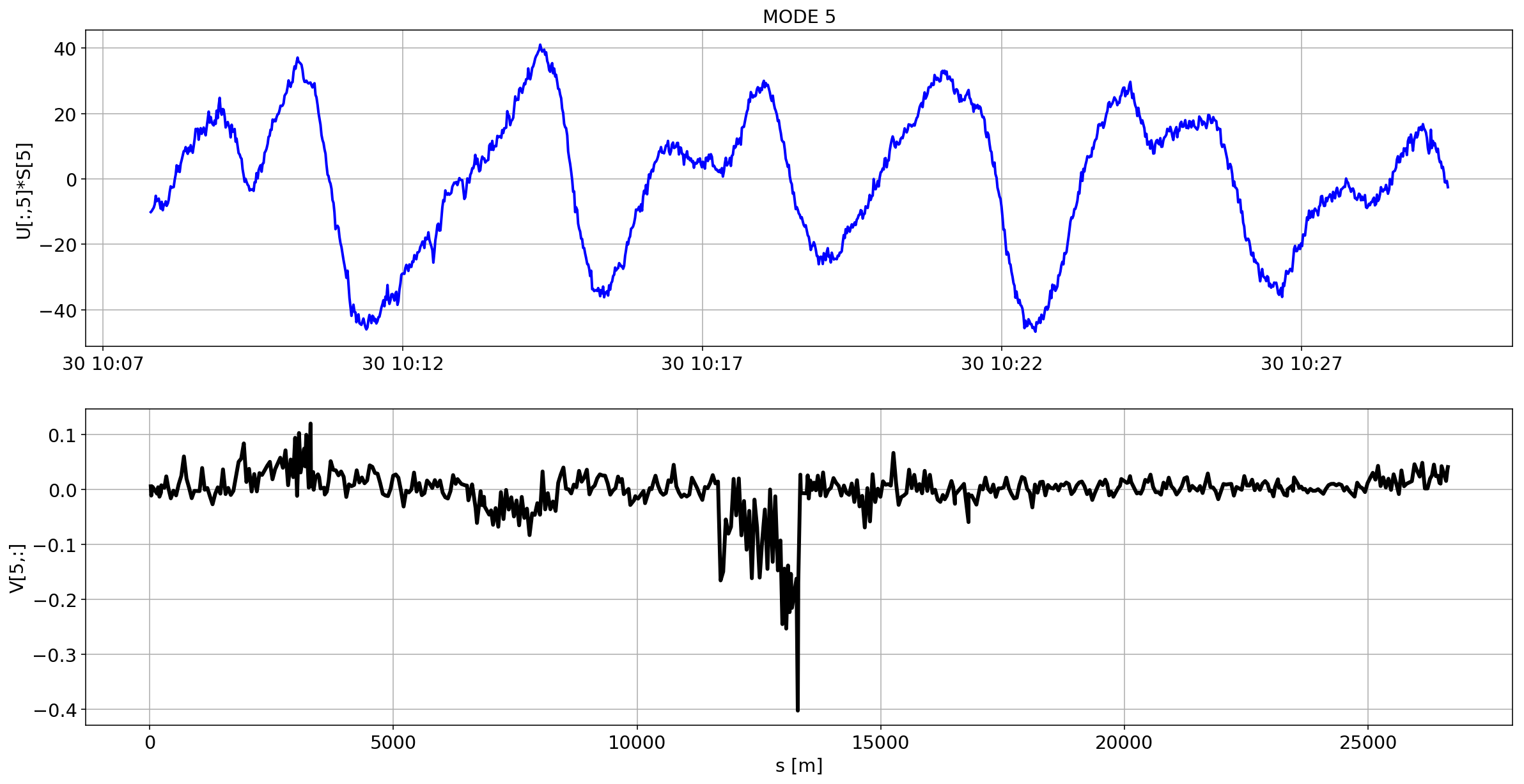
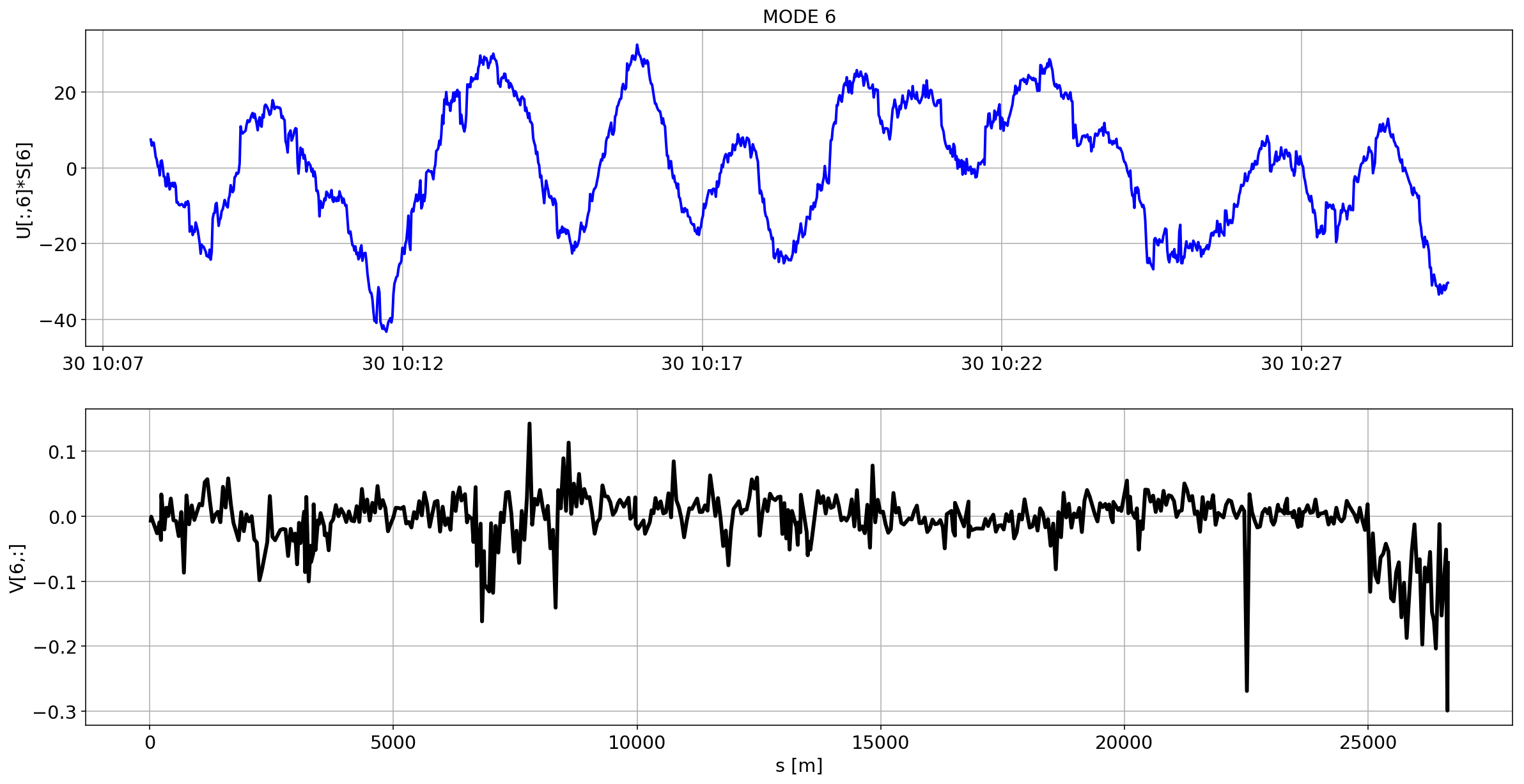
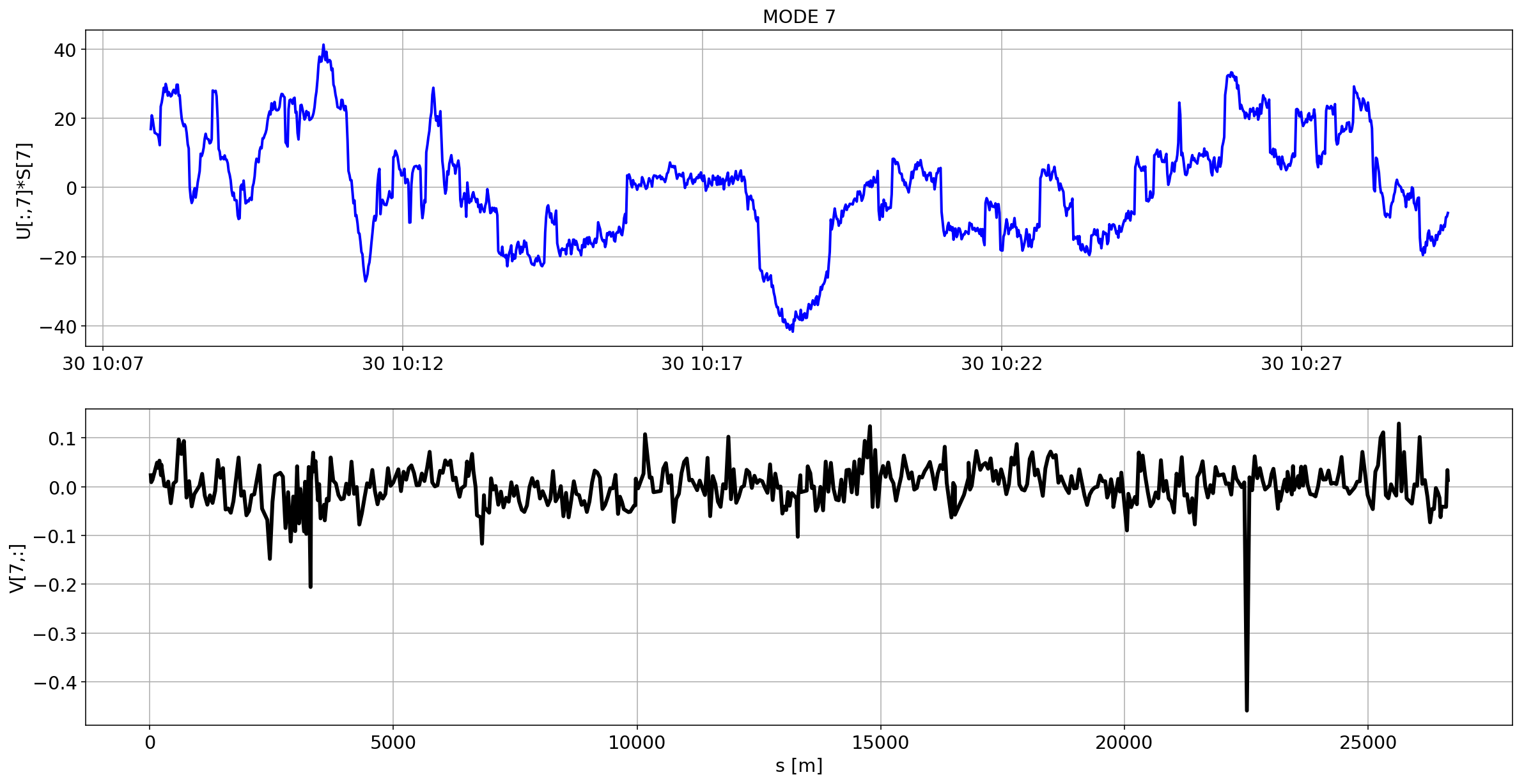
# we remove the mean orbit
aux=smallDF_B2-smallDF_B2.mean()
# we decompose the matrix
U,S,V=np.linalg.svd(aux.values,full_matrices=False)
plt.semilogy(S,'.-b')
#plt.xlim([0,40])
plt.xlabel('Singular Values number')
plt.ylabel('Singular Values amplitude');
plt.grid(True)
# sanity check
print(f'Sanity check: {np.allclose(aux.values,U * S @ V)}')
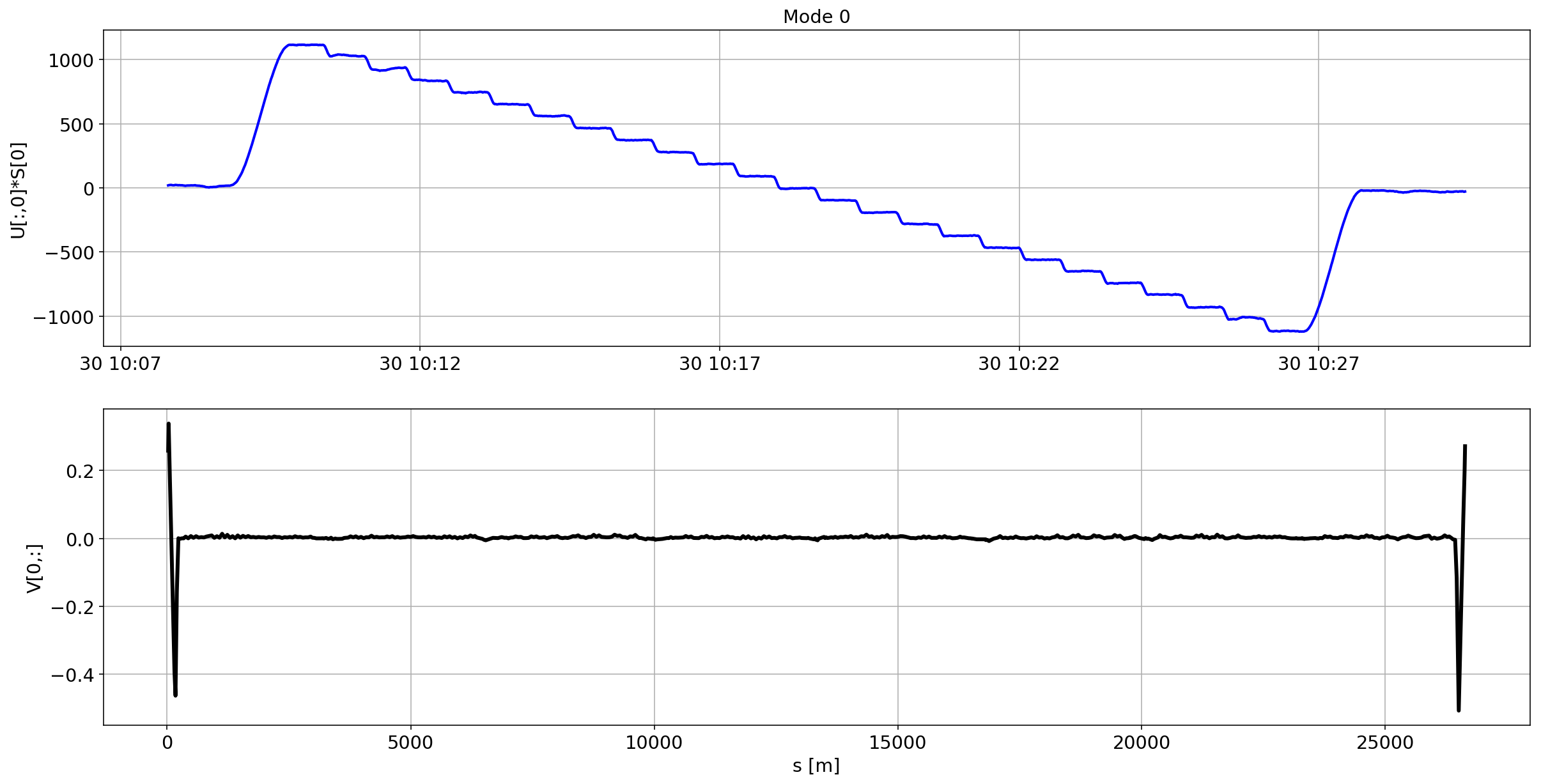
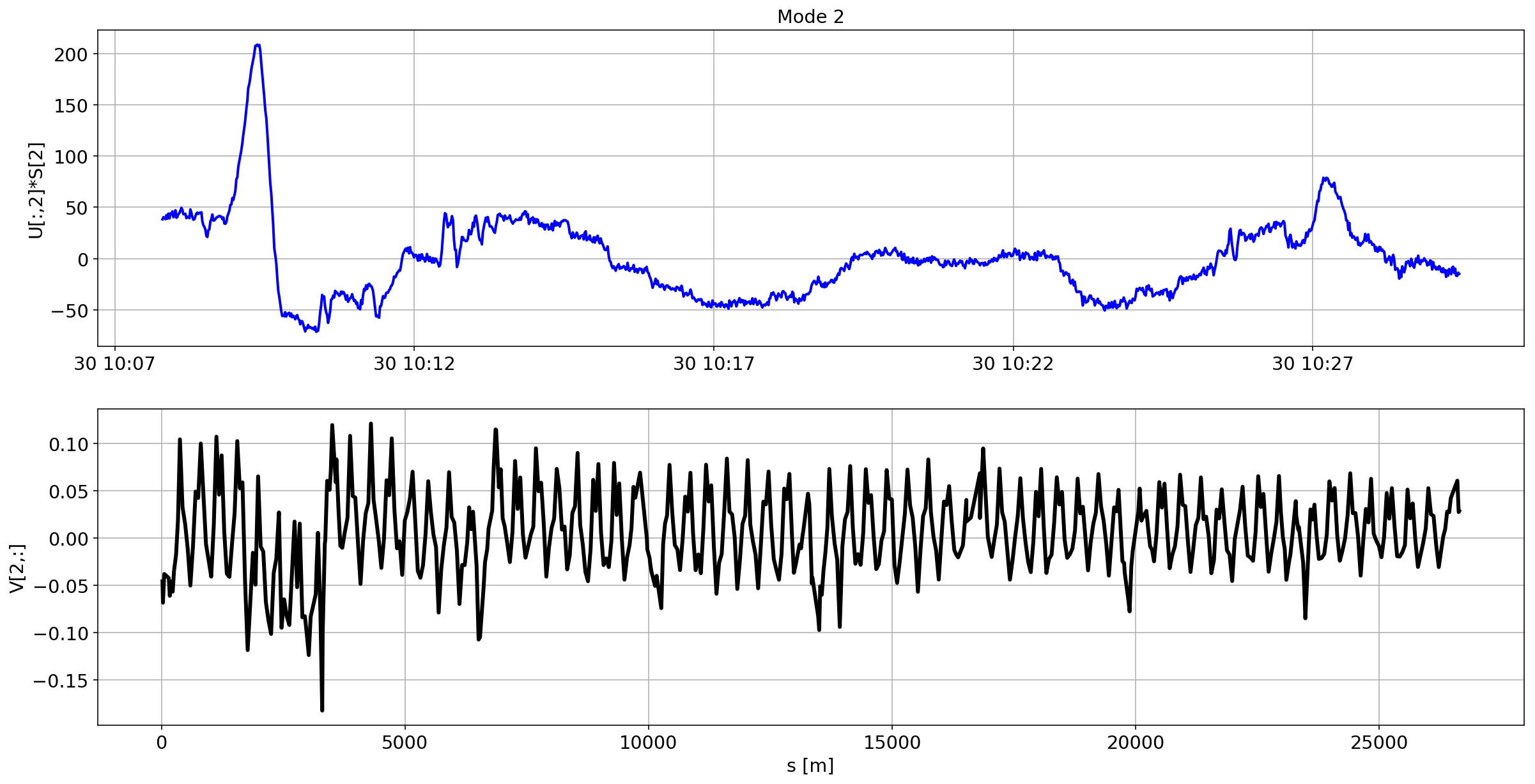
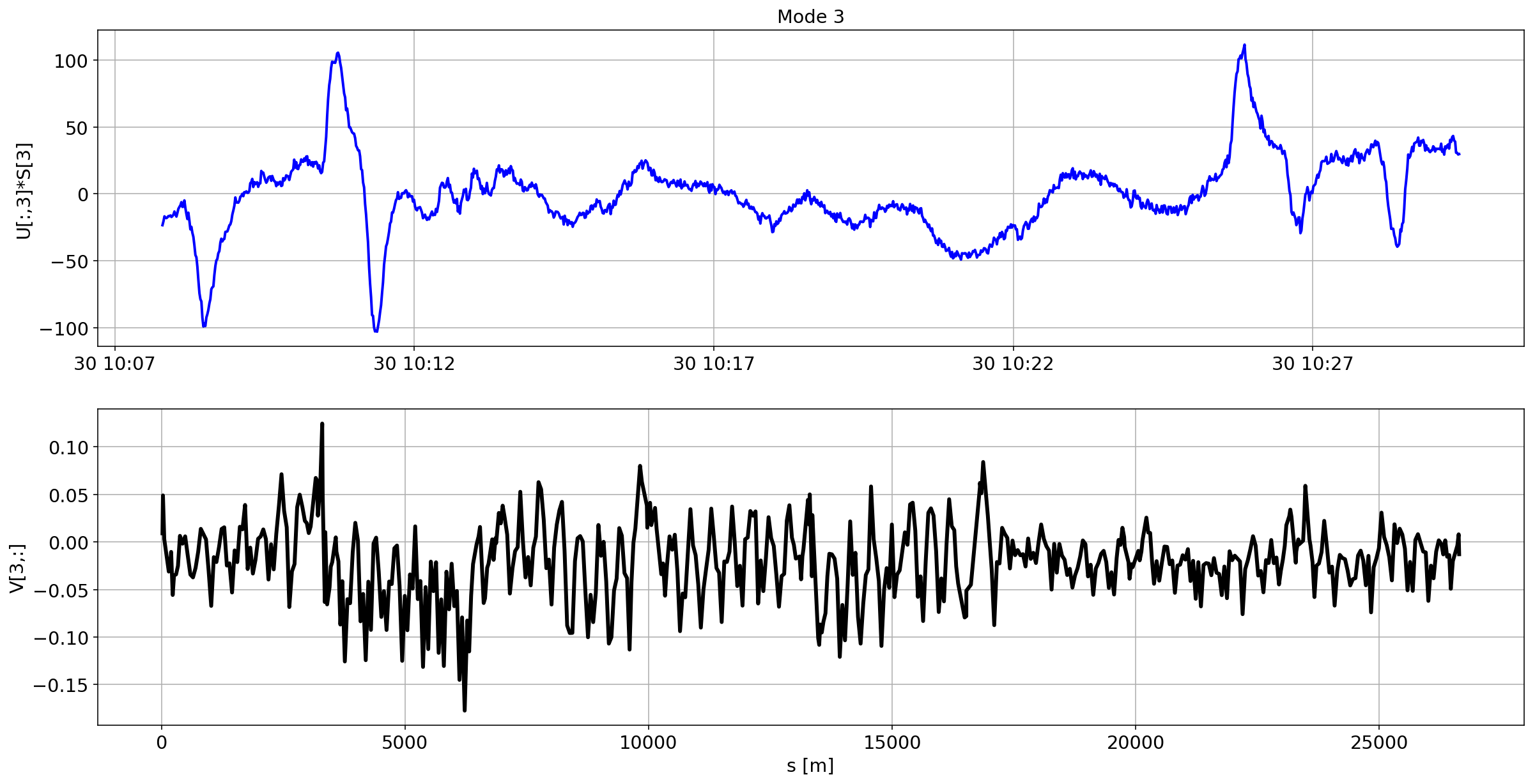
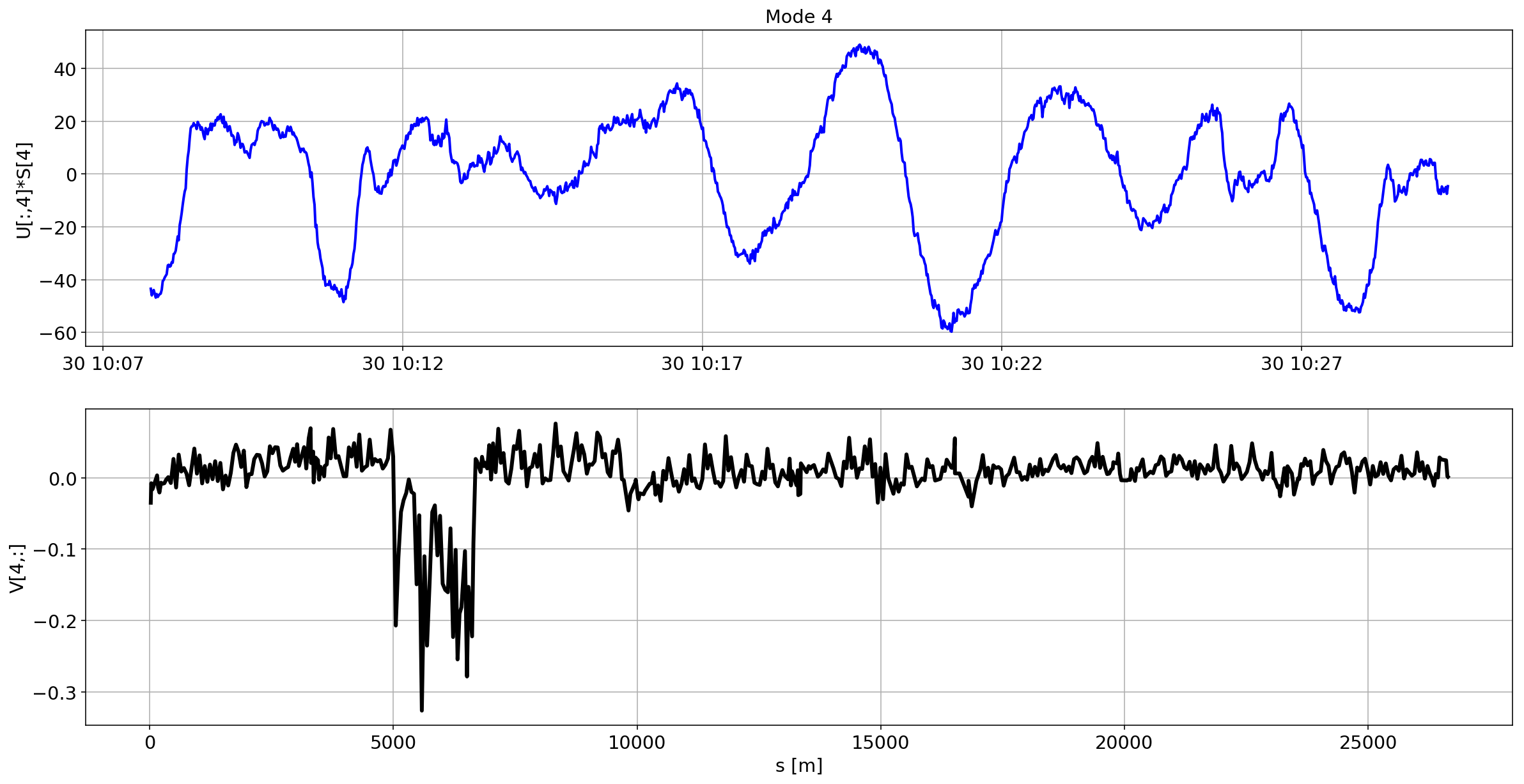
def plotMode_B2(mode=0):
plt.figure(figsize=(20,10))
plt.subplot(2,1,1)
plt.plot(aux.index,U[:,mode]*S[mode],'b',lw=2)
plt.grid(True)
plt.ylabel(f'U[:,{mode}]*S[{mode}]')
plt.title(f'Mode {mode}')
plt.subplot(2,1,2)
plt.plot(df_B2_H[df_B2_H['mask']==1]['s [m]'].values,V[mode,:],'k',lw=3)
plt.grid(True)
plt.ylabel(f'V[{mode},:]')
plt.xlabel('s [m]')
Sanity check: False
for i in range(10):
plotMode_B2(mode=i)
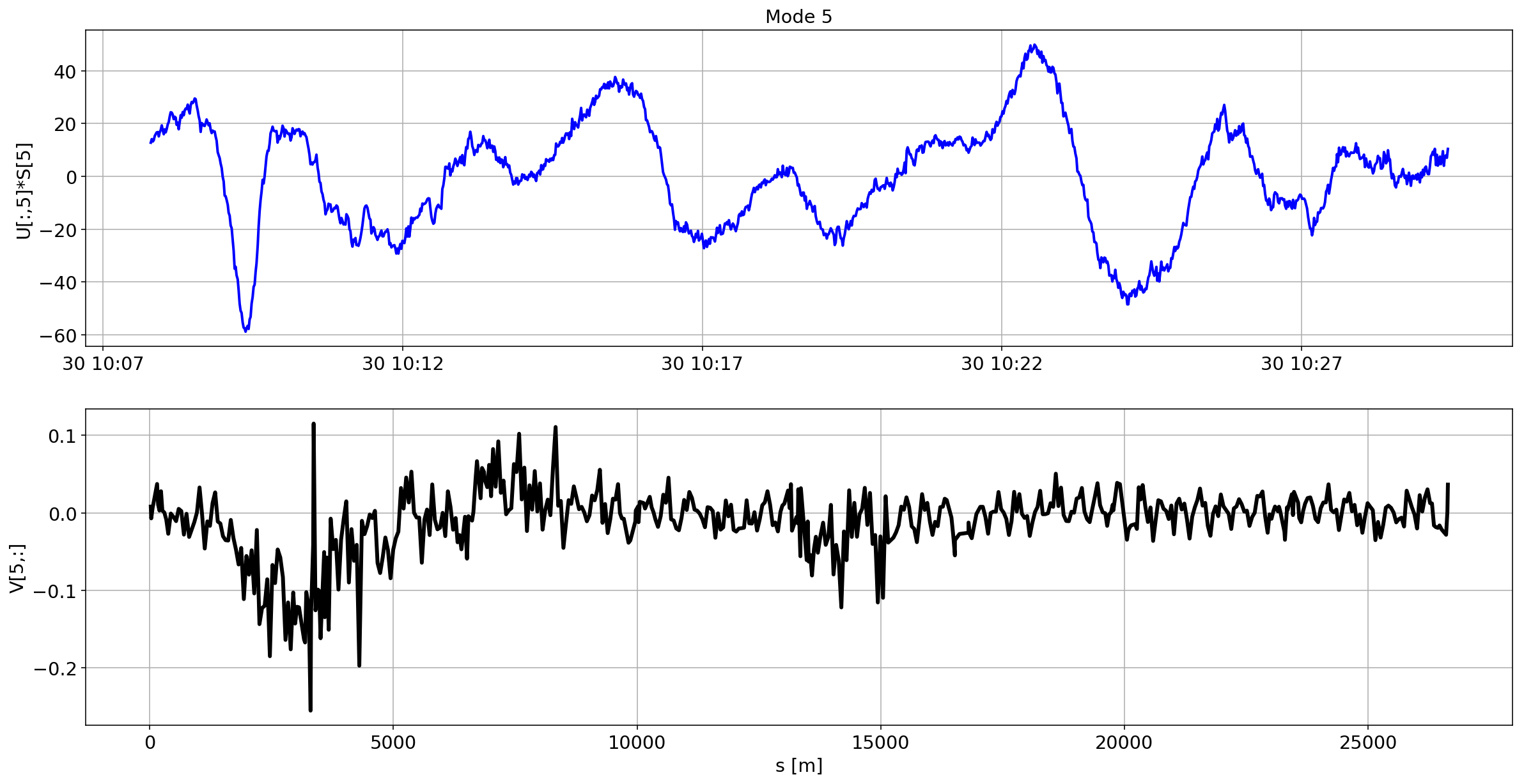
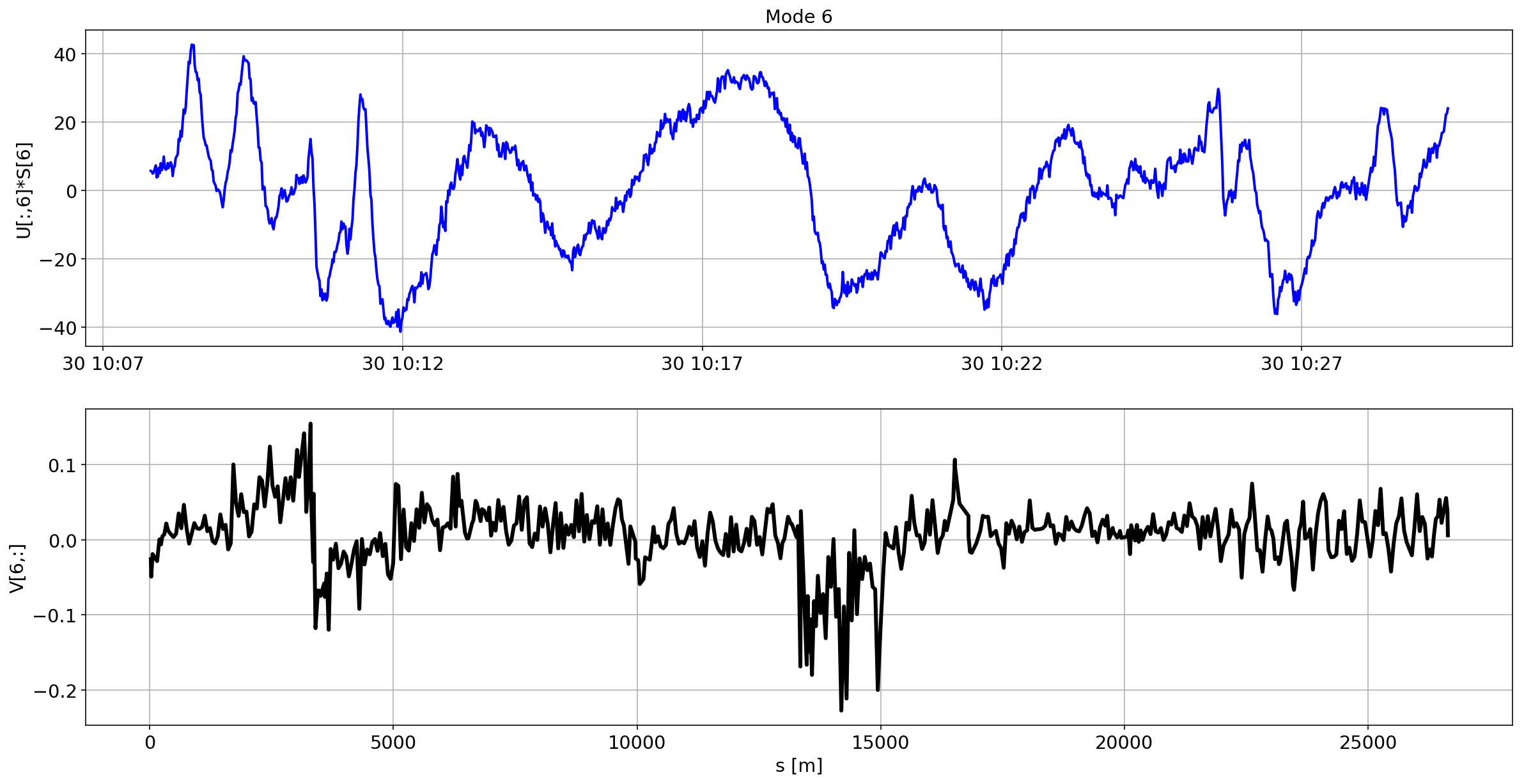
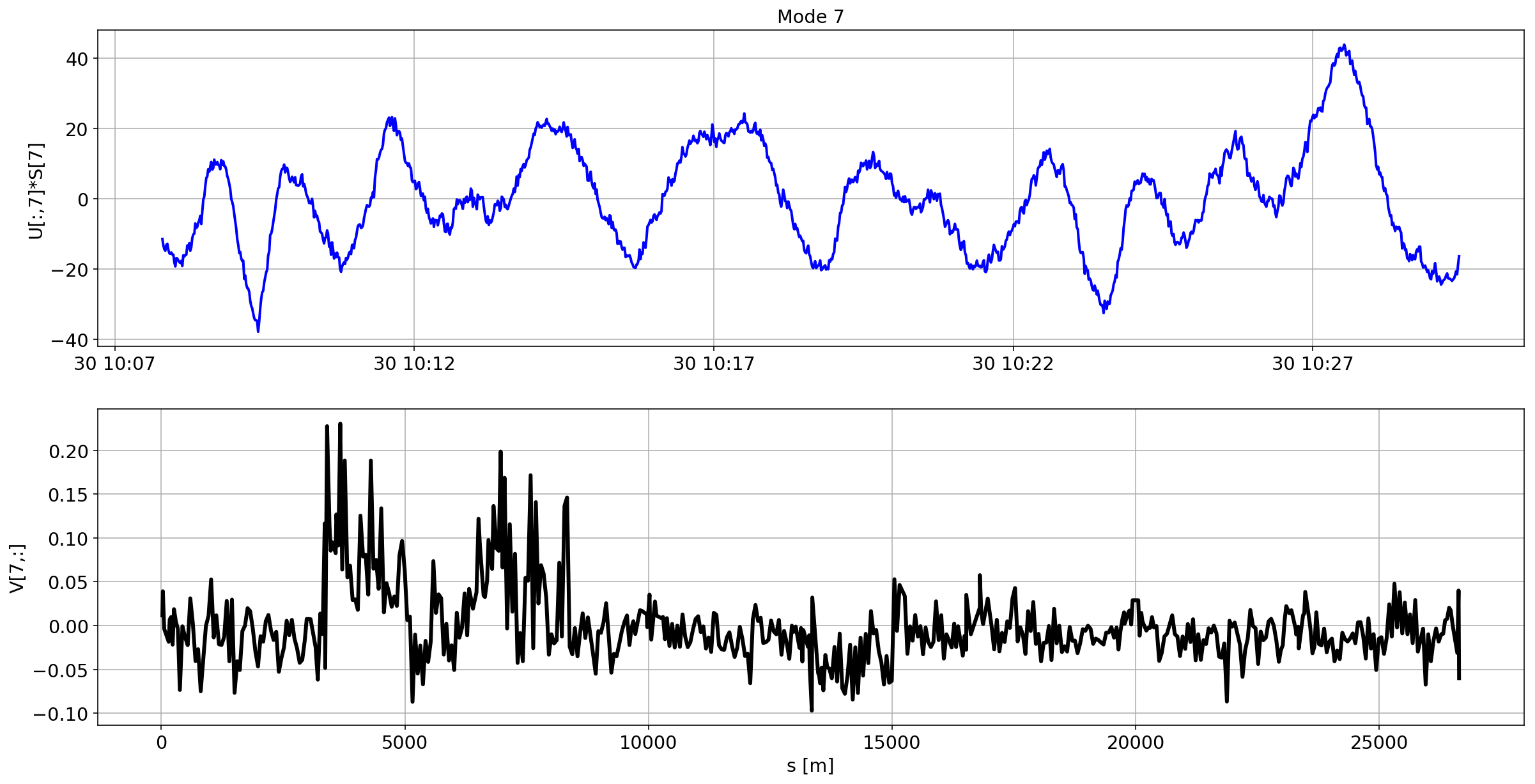
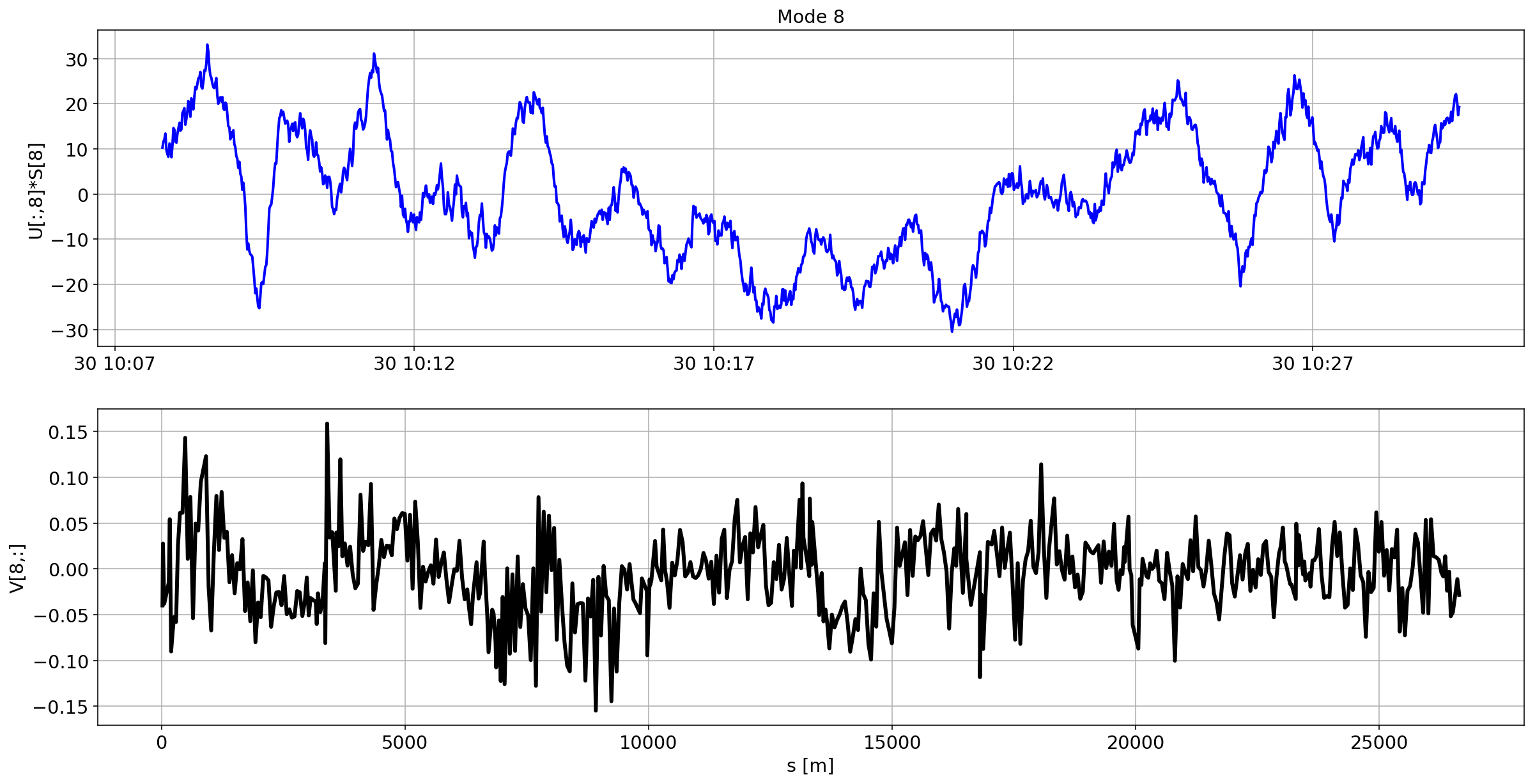
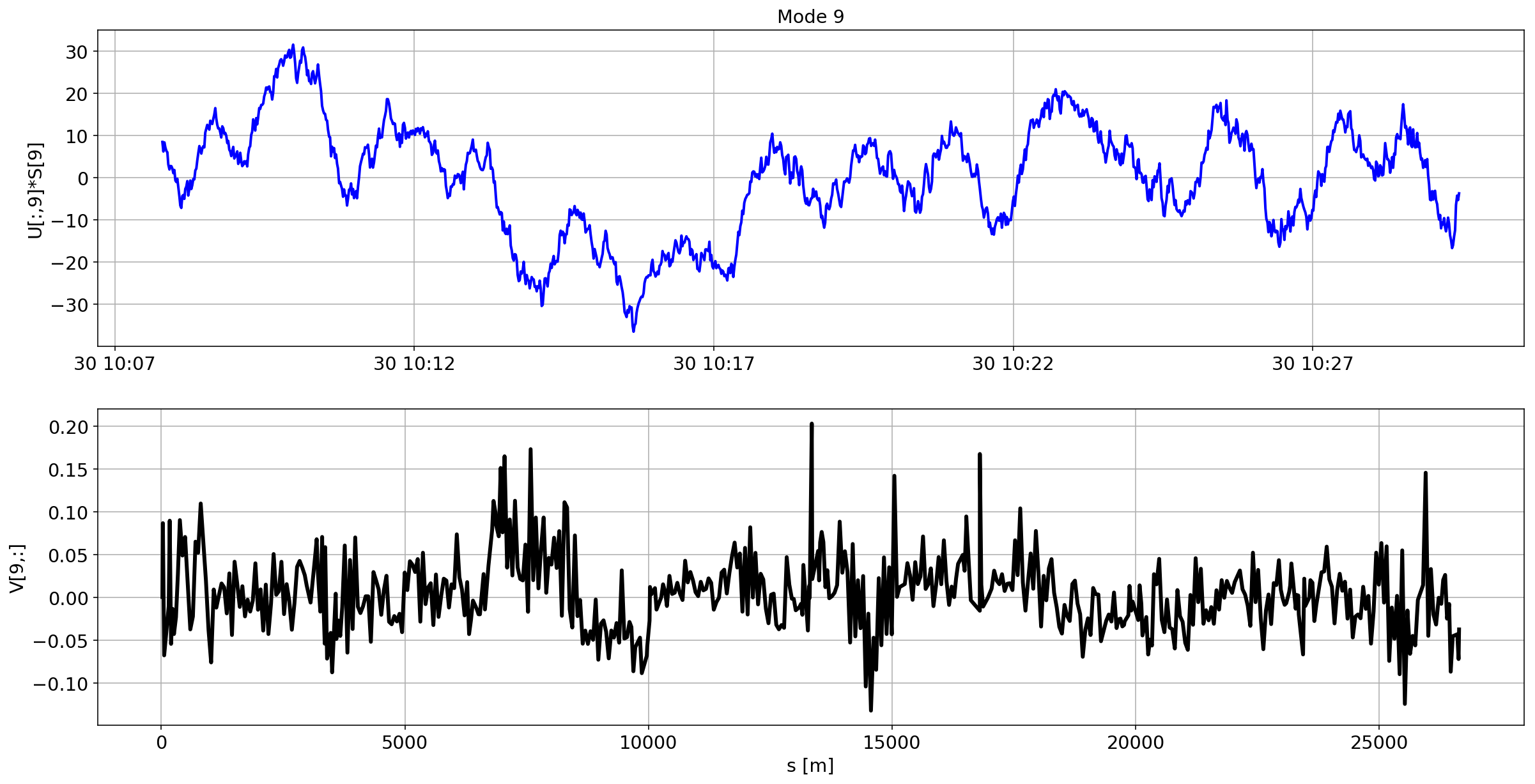
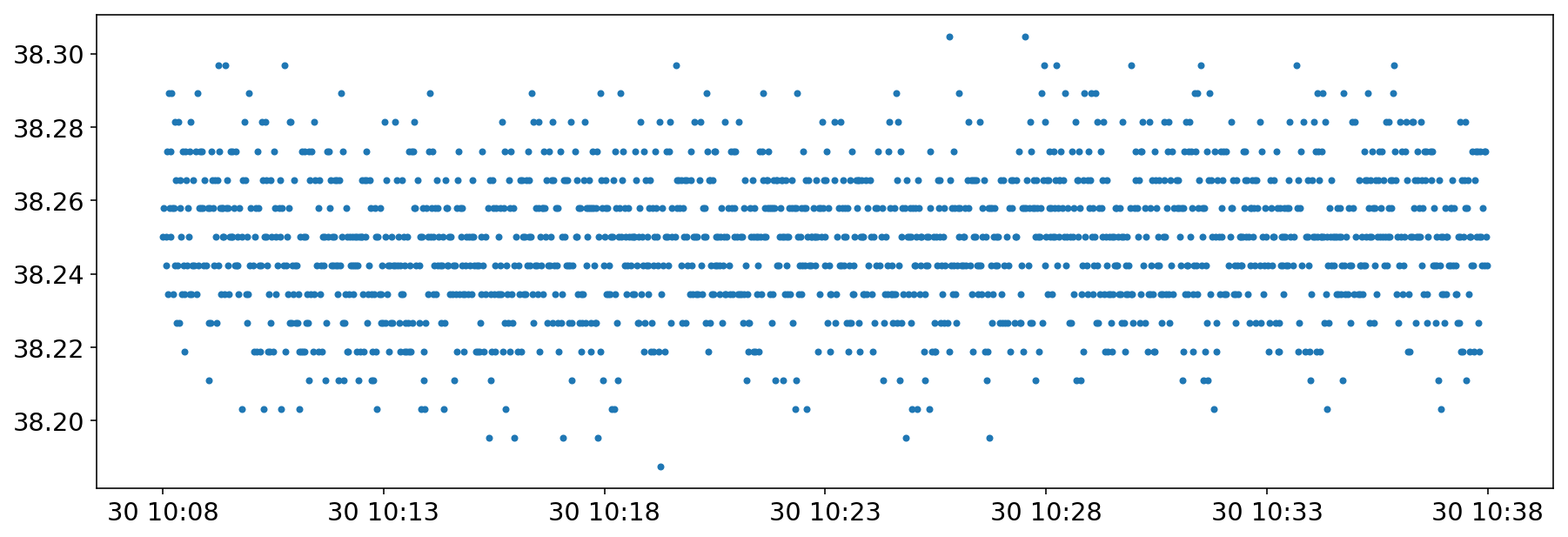
Tracking the temperature of the BPM
I started without much success (expert in holidays) to look to correlation between the temperature of the BPM and the beam oscillations.
cals.search('LHC.BPTUH.%.B1:%TEMP%')
['LHC.BPTUH.A4L1.B1:HW_TEMP_ADC',
'LHC.BPTUH.A4L1.B1:HW_TEMP_ANALOG',
'LHC.BPTUH.A4L1.B1:HW_TEMP_FPGA',
'LHC.BPTUH.A4L2.B1:HW_TEMP_ADC',
'LHC.BPTUH.A4L2.B1:HW_TEMP_ANALOG',
'LHC.BPTUH.A4L2.B1:HW_TEMP_FPGA',
'LHC.BPTUH.A4L5.B1:HW_TEMP_ADC',
'LHC.BPTUH.A4L5.B1:HW_TEMP_ANALOG',
'LHC.BPTUH.A4L5.B1:HW_TEMP_FPGA',
'LHC.BPTUH.A4L8.B1:HW_TEMP_ADC',
'LHC.BPTUH.A4L8.B1:HW_TEMP_ANALOG',
'LHC.BPTUH.A4L8.B1:HW_TEMP_FPGA',
'LHC.BPTUH.A4R6.B1:HW_TEMP_ADC',
'LHC.BPTUH.A4R6.B1:HW_TEMP_ANALOG',
'LHC.BPTUH.A4R6.B1:HW_TEMP_FPGA',
'LHC.BPTUH.C6L7.B1:HW_TEMP_ADC',
'LHC.BPTUH.C6L7.B1:HW_TEMP_ANALOG',
'LHC.BPTUH.C6L7.B1:HW_TEMP_FPGA']
plt.plot(importData.cals2pd('LHC.BPTUH.C6L7.B1:HW_TEMP_ANALOG',pd.Timestamp(2018,6,30,10,8),pd.Timestamp(2018,6,30,10,38)),'.')
[<matplotlib.lines.Line2D at 0x7fea87fb8ba8>]
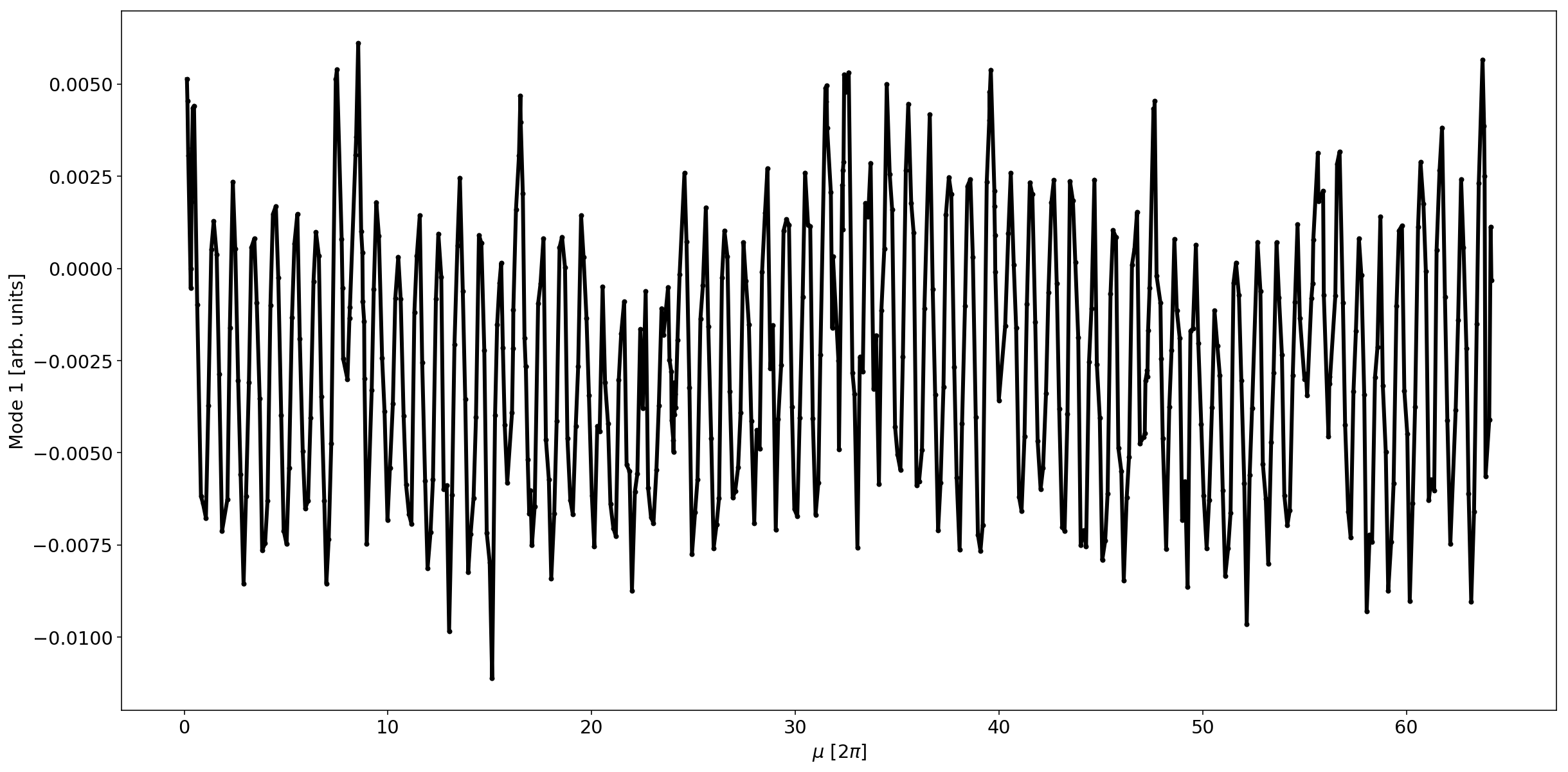
Localization of the error
betx=[]
mux=[]
for i in df_B2_H[df_B2_H['mask']==1]['s [m]'].values:
betx.append(b2DF[b2DF['s']==i].betx.values[0])
mux.append(b2DF[b2DF['s']==i].mux.values[0])
plt.figure(figsize=(20,10))
mode=1
plt.plot(mux,V[mode,:]/np.sqrt(betx),'.-k',lw=3)
#plt.xlim(0,10)
plt.xlabel('$\mu$ [2$\pi$]')
plt.ylabel(f'Mode {mode} [arb. units]');
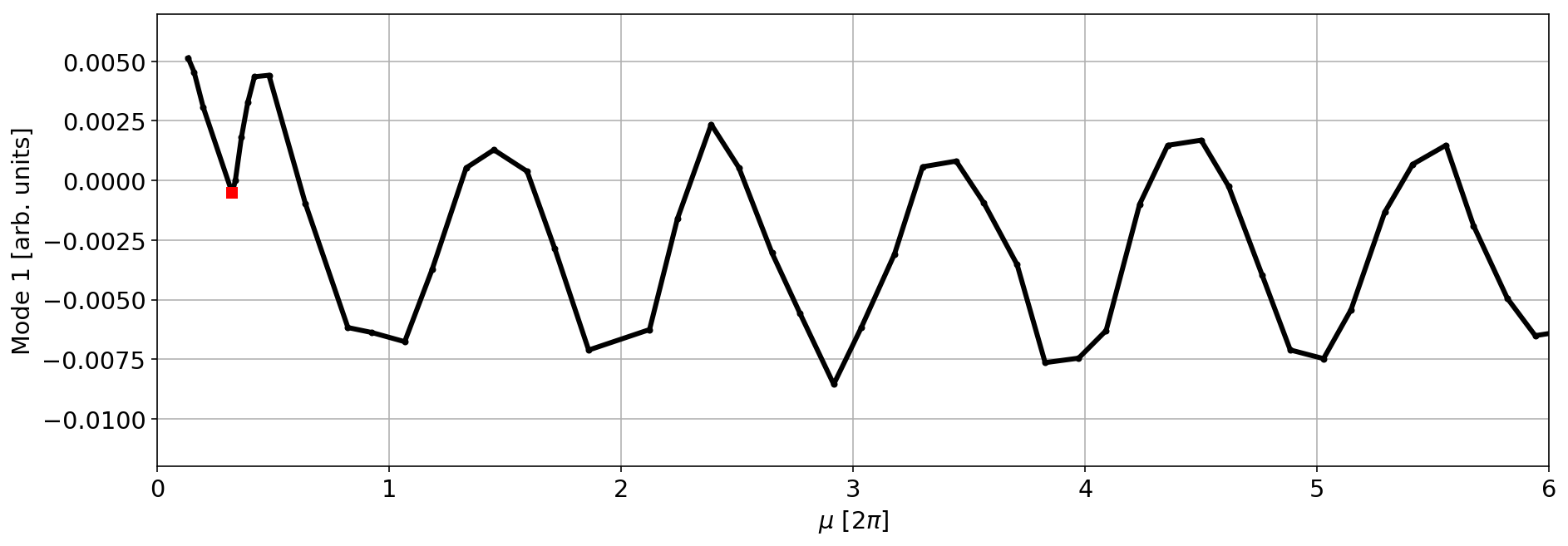
mode=1
plt.plot(mux,V[mode,:]/np.sqrt(betx),'.-k',lw=3)
myIndex=3
plt.plot(mux[myIndex],(V[mode,:]/np.sqrt(betx))[myIndex],'sr',lw=3)
plt.xlim(0,6)
plt.xlabel('$\mu$ [2$\pi$]')
plt.ylabel(f'Mode {mode} [arb. units]');
plt.grid(True)
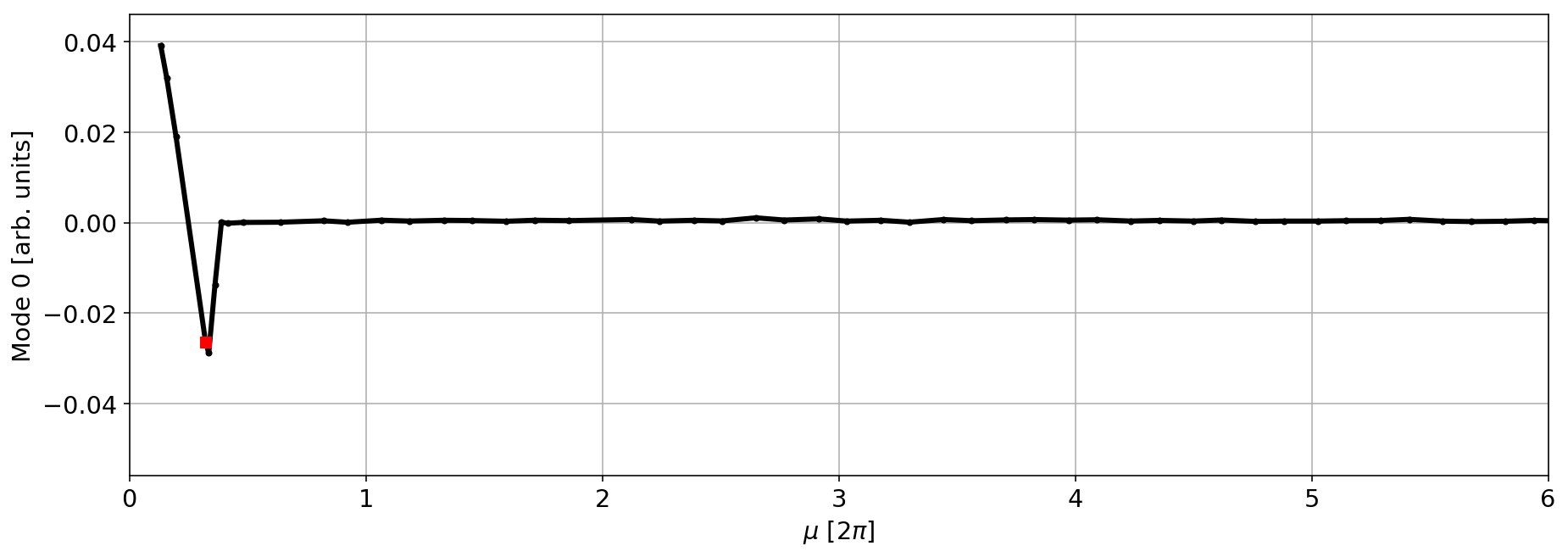
mode=0
plt.plot(mux,V[mode,:]/np.sqrt(betx),'.-k',lw=3)
myIndex=3
plt.plot(mux[myIndex],(V[mode,:]/np.sqrt(betx))[myIndex],'sr',lw=3)
plt.xlim(0,6)
plt.xlabel('$\mu$ [2$\pi$]')
plt.ylabel(f'Mode {mode} [arb. units]');
plt.grid(True)
SVD filtering
# we remove the mean orbit
aux=smallDF_B1-smallDF_B1.mean()
# we decompose the matrix
U,S,V=np.linalg.svd(aux.values,full_matrices=False)
# sanity check
print(f'Sanity check: {np.allclose(aux.values,U * S @ V)}')
Sanity check: True
# taking only the first mode
plt.figure(figsize=(20,10))
myS=S.copy()
myS[1:]=0
plt.pcolormesh(U * myS @ V *1e-3)
ax=plt.colorbar()
plt.xlabel('BPM')
plt.ylabel('time [arb. units]')
ax.set_label('B1 H orbit [mm]')

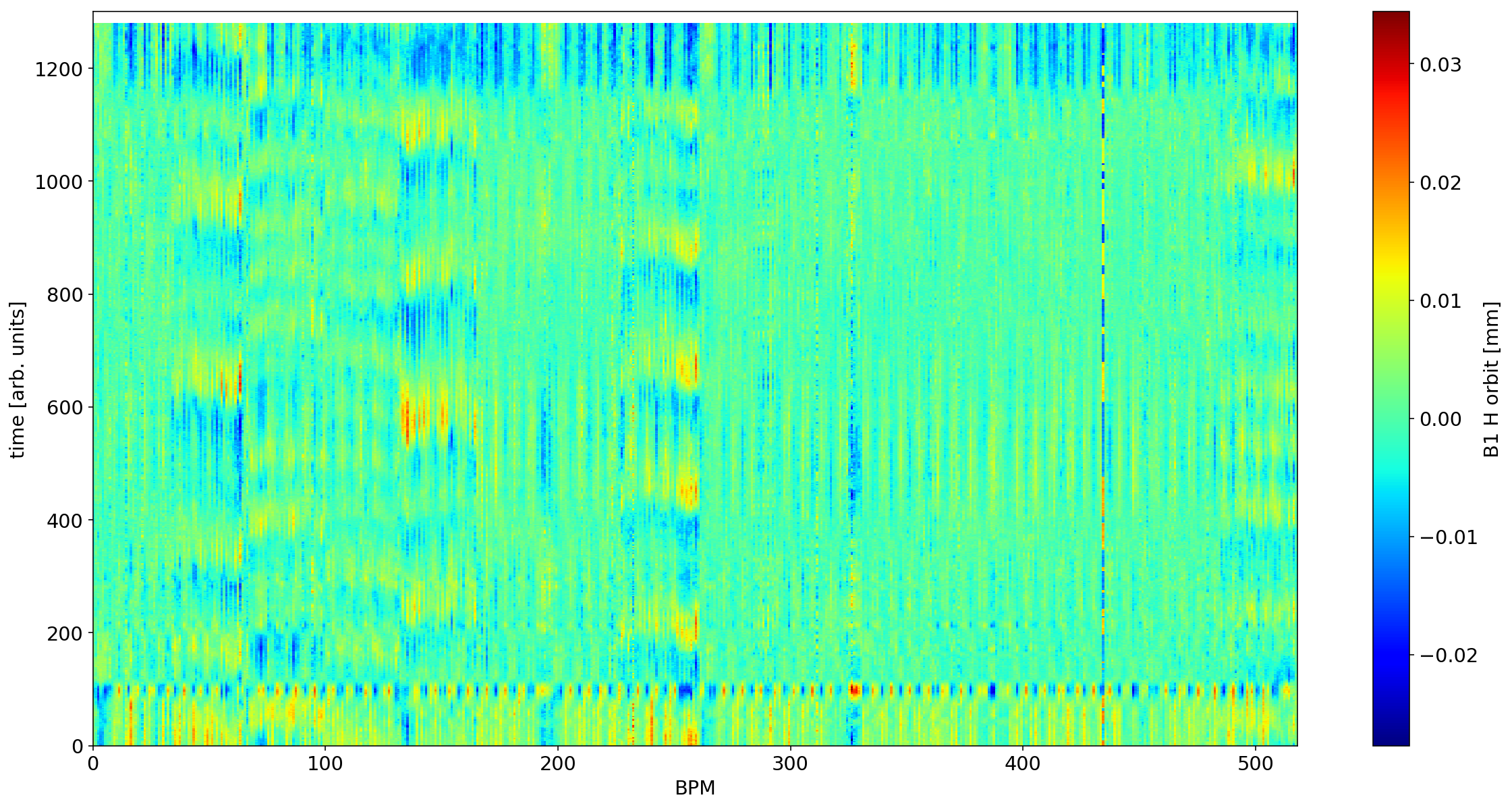
# removing the first singular value
plt.figure(figsize=(20,10))
myS=S.copy()
myS[0]=0
plt.pcolormesh(U * myS @ V *1e-3)
ax=plt.colorbar()
plt.xlabel('BPM')
plt.ylabel('time [arb. units]')
ax.set_label('B1 H orbit [mm]')
# we remove the mean orbit
aux=smallDF_B2-smallDF_B2.mean()
# we decompose the matrix
U,S,V=np.linalg.svd(aux.values,full_matrices=False)
# sanity check
print(f'Sanity check: {np.allclose(aux.values,U * S @ V)}')
Sanity check: True
# taking only the first singular value
plt.figure(figsize=(20,10))
myS=S.copy()
myS[1:]=0
plt.pcolormesh(U * myS @ V *1e-3)
ax=plt.colorbar()
plt.xlabel('BPM')
plt.ylabel('time [arb. units]')
ax.set_label('B2 H orbit [mm]')

# removing the first singular value
plt.figure(figsize=(20,10))
myS=S.copy()
myS[0]=0
plt.pcolormesh(U * myS @ V *1e-3)
ax=plt.colorbar()
plt.xlabel('BPM')
plt.ylabel('time [arb. units]')
ax.set_label('B2 H orbit [mm]')
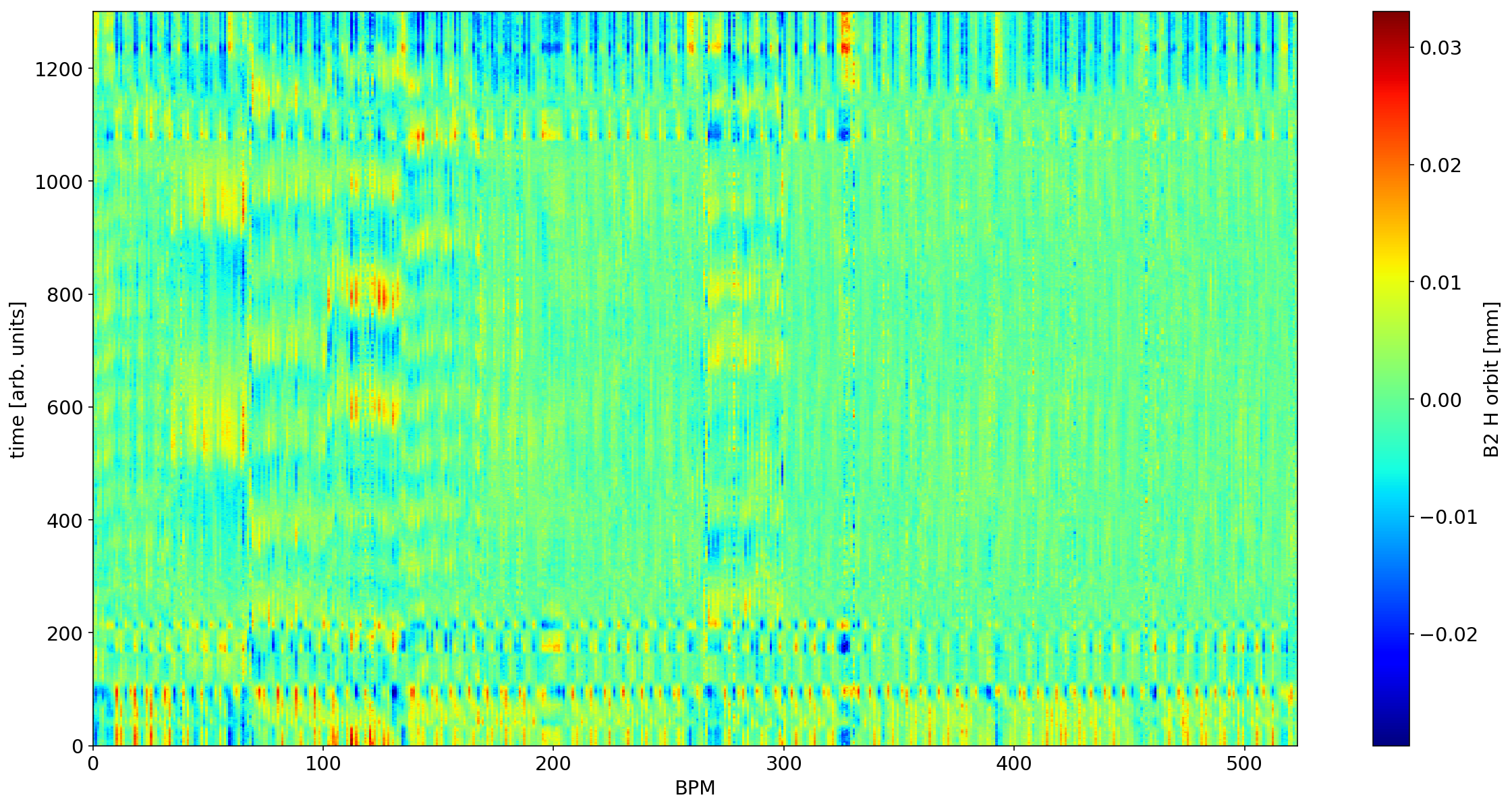
Conclusions
-
The preliminary analysis of the orbit stability during FILL 6868 is compatible with histeresis effect of the separation bumps (see mode=1).
-
After normalization, the mode=1 in B2 seems to reveal a spurious kick within the separation bump.
-
The analysis was perfomed only for the first scan (H-plane scan in IP1 and H-plane).
-
Some families of BPM show oscillations of their electrical center (clearly non physical).
-
The analysis spotted also a very reduced number of problematic BPM (in particular one in B1/H).
-
The SVD analysis can combine also additional signal signals (e.g., V plane, DOROS, thermal sensor of BPM crate) to improve the reading.
Optics during the vdm scan
importData.LHCCals2pd(['HX:OPTID'],myFill,beamModeList=['INJPROT','INJPHYS','PRERAMP','RAMP','FLATTOP','ADJUST','STABLE'],beamMode_column=True)
| HX:OPTID | mode | |
|---|---|---|
| 2018-06-30 09:13:12.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:14:12.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:15:12.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:16:32.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:18:12.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:19:52.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:21:32.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:22:52.046000004+00:00 | 3215.0 | RAMP |
| 2018-06-30 09:24:02.046000004+00:00 | 3226.0 | RAMP |
| 2018-06-30 09:25:02.046000004+00:00 | 3220.0 | RAMP |
| 2018-06-30 09:26:02.046000004+00:00 | 3223.0 | RAMP |
| 2018-06-30 09:27:02.046000004+00:00 | 3222.0 | RAMP |
| 2018-06-30 09:28:02.046000004+00:00 | 3221.0 | RAMP |
| 2018-06-30 09:29:02.046000004+00:00 | 3225.0 | RAMP |
| 2018-06-30 09:31:32.046000004+00:00 | 3225.0 | RAMP |
| 2018-06-30 09:33:22.046000004+00:00 | 3225.0 | RAMP |
| 2018-06-30 09:38:09.472000122+00:00 | 3225.0 | FLATTOP |
| 2018-06-30 09:38:29.472000122+00:00 | 3224.0 | FLATTOP |
| 2018-06-30 09:41:07.421999931+00:00 | 3224.0 | ADJUST |
| 2018-06-30 09:42:27.421999931+00:00 | 3224.0 | ADJUST |
| 2018-06-30 09:43:47.421999931+00:00 | 3224.0 | ADJUST |
| 2018-06-30 09:45:55.582999945+00:00 | 3224.0 | ADJUST |
Optics computation
from cpymad.madx import Madx
from matplotlib import pyplot as plt
myString='''
! Sequence definition
Option, -echo,warn,-info;
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/const_for_aperture.madx";
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/lhc_as-built.seq";
! Aperture definition
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/aperture_as-built.b1.madx";
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/aperture_as-built.b2.madx";
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/aper_tol_as-built.b1.madx";
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/aper_tol_as-built.b2.madx";
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/exp_pipe_model_after_LS1.madx";
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/aperture/exp_pipe_install_after_LS1.madx";
! Beam definition
beam, sequence=lhcb1, bv= 1,
particle=proton, charge=1, mass=0.938272046,
energy= 6500, npart=1.2e11,kbunch=2748,
ex=3.608738638461539e-10,ey=3.608738638461539e-10;
beam, sequence=lhcb2, bv=-1,
particle=proton, charge=1, mass=0.938272046,
energy= 6500, npart=1.2e11,kbunch=2748,
ex=3.608738638461539e-10,ey=3.608738638461539e-10;
! Strength definition
call, file="/eos/project/a/abpdata/lhc/optics/runII/2015/opt_19200_19000_19200_24000_coll.madx";
nrj=beam%lhcb1->pc/beam%lhcb1->charge;
! Cycle
!seqedit,sequence=lhcb1;flatten;cycle,start=s.ds.l1.b1;endedit;
!seqedit,sequence=lhcb2;flatten;cycle,start=s.ds.l1.b2;endedit;
! Post Strength
! Twiss before customized knobs
set,format="22.15e";
select,flag=twiss,clear;
select,flag=twiss, pattern="IP.$",column=name,s,betx,bety,alfx,alfy,dx,dpx,dy,dpy,mux,muy,x,y,px,py;
use, sequence=lhcb1;
twiss, table=twiss_ipb1_flat;
use, sequence=lhcb2;
twiss, table=twiss_ipb2_flat;
'''
myMad = Madx()
++++++++++++++++++++++++++++++++++++++++++++
+ MAD-X 5.04.02 (64 bit, Linux) +
+ Support: mad@cern.ch, http://cern.ch/mad +
+ Release date: 2018.10.03 +
+ Execution date: 2019.07.03 09:53:26 +
++++++++++++++++++++++++++++++++++++++++++++
myMad.input(myString)
++++++ warning: ignored: attempt to redefine constant: l.mbas2
++++++ warning: ignored: attempt to redefine constant: l.mbaw
++++++ warning: ignored: attempt to redefine constant: l.mbcs2
++++++ warning: ignored: attempt to redefine constant: l.mbls2
++++++ warning: ignored: attempt to redefine constant: l.mblw
++++++ warning: ignored: attempt to redefine constant: l.mbwmd
++++++ warning: ignored: attempt to redefine constant: l.mbxwh
++++++ warning: ignored: attempt to redefine constant: l.mbxws
++++++ warning: ignored: attempt to redefine constant: l.mbxwt
++++++ warning: implicit element re-definition ignored: ip1
++++++ warning: implicit element re-definition ignored: mbas2.1r1
++++++ warning: implicit element re-definition ignored: tas.1r1
++++++ warning: implicit element re-definition ignored: bpmwk.1r1
++++++ warning: implicit element re-definition ignored: mqxa.1r1
++++++ warning: implicit element re-definition ignored: mcbxh.1r1
++++++ warning: implicit element re-definition ignored: mcbxv.1r1
++++++ warning: implicit element re-definition ignored: mqxb.a2r1
++++++ warning: implicit element re-definition ignored: mcbxh.2r1
++++++ warning: implicit element re-definition ignored: mcbxv.2r1
++++++ warning: implicit element re-definition ignored: mqxb.b2r1
++++++ warning: implicit element re-definition ignored: tasb.3r1
++++++ warning: implicit element re-definition ignored: mqsx.3r1
++++++ warning: implicit element re-definition ignored: mqxa.3r1
++++++ warning: implicit element re-definition ignored: mcbxh.3r1
++++++ warning: implicit element re-definition ignored: mcbxv.3r1
++++++ warning: implicit element re-definition ignored: mcsx.3r1
++++++ warning: implicit element re-definition ignored: mctx.3r1
++++++ warning: implicit element re-definition ignored: mcosx.3r1
++++++ warning: implicit element re-definition ignored: mcox.3r1
++++++ warning: implicit element re-definition ignored: mcssx.3r1
++++++ warning: implicit element re-definition ignored: dfbxb.3r1
++++++ warning: implicit element re-definition ignored: mbxw.a4r1
++++++ warning: implicit element re-definition ignored: mbxw.b4r1
++++++ warning: implicit element re-definition ignored: mbxw.c4r1
++++++ warning: implicit element re-definition ignored: mbxw.d4r1
++++++ warning: implicit element re-definition ignored: mbxw.e4r1
++++++ warning: implicit element re-definition ignored: mbxw.f4r1
++++++ warning: implicit element re-definition ignored: x1fcr.4r1
++++++ warning: implicit element re-definition ignored: brana.4r1
++++++ warning: implicit element re-definition ignored: x1zdc.a4r1
++++++ warning: implicit element re-definition ignored: tanar.4r1
++++++ warning: implicit element re-definition ignored: x2zdc.4l2
++++++ warning: implicit element re-definition ignored: branb.4l2
++++++ warning: implicit element re-definition ignored: btvst.a4l2
++++++ warning: implicit element re-definition ignored: tcdd.4l2
++++++ warning: implicit element re-definition ignored: mbx.4l2
++++++ warning: implicit element re-definition ignored: dfbxc.3l2
++++++ warning: implicit element re-definition ignored: mcosx.3l2
++++++ warning: implicit element re-definition ignored: mcox.3l2
++++++ warning: implicit element re-definition ignored: mcssx.3l2
++++++ warning: implicit element re-definition ignored: mcbxh.3l2
++++++ warning: implicit element re-definition ignored: mcbxv.3l2
++++++ warning: implicit element re-definition ignored: mcsx.3l2
++++++ warning: implicit element re-definition ignored: mctx.3l2
++++++ warning: implicit element re-definition ignored: mqxa.3l2
++++++ warning: implicit element re-definition ignored: mqsx.3l2
++++++ warning: implicit element re-definition ignored: mqxb.b2l2
++++++ warning: implicit element re-definition ignored: mcbxh.2l2
++++++ warning: implicit element re-definition ignored: mcbxv.2l2
++++++ warning: implicit element re-definition ignored: mqxb.a2l2
++++++ warning: implicit element re-definition ignored: mcbxh.1l2
++++++ warning: implicit element re-definition ignored: mcbxv.1l2
++++++ warning: implicit element re-definition ignored: mqxa.1l2
++++++ warning: implicit element re-definition ignored: mbxwt.1l2
++++++ warning: implicit element re-definition ignored: mbwmd.1l2
++++++ warning: implicit element re-definition ignored: mbls2.1l2
++++++ warning: implicit element re-definition ignored: ip2
++++++ warning: implicit element re-definition ignored: mbls2.1r2
++++++ warning: implicit element re-definition ignored: mbaw.1r2
++++++ warning: implicit element re-definition ignored: mbxwt.1r2
++++++ warning: implicit element re-definition ignored: mqxa.1r2
++++++ warning: implicit element re-definition ignored: mcbxh.1r2
++++++ warning: implicit element re-definition ignored: mcbxv.1r2
++++++ warning: implicit element re-definition ignored: mqxb.a2r2
++++++ warning: implicit element re-definition ignored: mcbxh.2r2
++++++ warning: implicit element re-definition ignored: mcbxv.2r2
++++++ warning: implicit element re-definition ignored: mqxb.b2r2
++++++ warning: implicit element re-definition ignored: mqsx.3r2
++++++ warning: implicit element re-definition ignored: mqxa.3r2
++++++ warning: implicit element re-definition ignored: mcbxh.3r2
++++++ warning: implicit element re-definition ignored: mcbxv.3r2
++++++ warning: implicit element re-definition ignored: mcsx.3r2
++++++ warning: implicit element re-definition ignored: mctx.3r2
++++++ warning: implicit element re-definition ignored: mcosx.3r2
++++++ warning: implicit element re-definition ignored: mcox.3r2
++++++ warning: implicit element re-definition ignored: mcssx.3r2
++++++ warning: implicit element re-definition ignored: dfbxd.3r2
++++++ warning: implicit element re-definition ignored: mbx.4r2
++++++ warning: implicit element re-definition ignored: tclia.4r2
++++++ warning: implicit element re-definition ignored: branb.4r2
++++++ warning: implicit element re-definition ignored: x2zdc.4r2
++++++ warning: implicit element re-definition ignored: ip3
++++++ warning: implicit element re-definition ignored: ip4
++++++ warning: implicit element re-definition ignored: tanc.4l5
++++++ warning: implicit element re-definition ignored: x5zdc.b4l5
++++++ warning: implicit element re-definition ignored: brana.4l5
++++++ warning: implicit element re-definition ignored: x5zdc.a4l5
++++++ warning: implicit element re-definition ignored: x5fcb.a4l5
++++++ warning: implicit element re-definition ignored: x5fca.b4l5
++++++ warning: implicit element re-definition ignored: mbxw.f4l5
++++++ warning: implicit element re-definition ignored: mbxw.e4l5
++++++ warning: implicit element re-definition ignored: mbxw.d4l5
++++++ warning: implicit element re-definition ignored: mbxw.c4l5
++++++ warning: implicit element re-definition ignored: mbxw.b4l5
++++++ warning: implicit element re-definition ignored: mbxw.a4l5
++++++ warning: implicit element re-definition ignored: x5fca.a4l5
++++++ warning: implicit element re-definition ignored: dfbxe.3l5
++++++ warning: implicit element re-definition ignored: mcosx.3l5
++++++ warning: implicit element re-definition ignored: mcox.3l5
++++++ warning: implicit element re-definition ignored: mcssx.3l5
++++++ warning: implicit element re-definition ignored: mcbxh.3l5
++++++ warning: implicit element re-definition ignored: mcbxv.3l5
++++++ warning: implicit element re-definition ignored: mcsx.3l5
++++++ warning: implicit element re-definition ignored: mctx.3l5
++++++ warning: implicit element re-definition ignored: mqxa.3l5
++++++ warning: implicit element re-definition ignored: mqsx.3l5
++++++ warning: implicit element re-definition ignored: tasb.3l5
++++++ warning: implicit element re-definition ignored: mqxb.b2l5
++++++ warning: implicit element re-definition ignored: mcbxh.2l5
++++++ warning: implicit element re-definition ignored: mcbxv.2l5
++++++ warning: implicit element re-definition ignored: mqxb.a2l5
++++++ warning: implicit element re-definition ignored: mcbxh.1l5
++++++ warning: implicit element re-definition ignored: mcbxv.1l5
++++++ warning: implicit element re-definition ignored: mqxa.1l5
++++++ warning: implicit element re-definition ignored: bpmwk.1l5
++++++ warning: implicit element re-definition ignored: tas.1l5
++++++ warning: implicit element re-definition ignored: mbcs2.1l5
++++++ warning: implicit element re-definition ignored: ip5
++++++ warning: implicit element re-definition ignored: mbcs2.1r5
++++++ warning: implicit element re-definition ignored: tas.1r5
++++++ warning: implicit element re-definition ignored: bpmwk.1r5
++++++ warning: implicit element re-definition ignored: mqxa.1r5
++++++ warning: implicit element re-definition ignored: mcbxh.1r5
++++++ warning: implicit element re-definition ignored: mcbxv.1r5
++++++ warning: implicit element re-definition ignored: mqxb.a2r5
++++++ warning: implicit element re-definition ignored: mcbxh.2r5
++++++ warning: implicit element re-definition ignored: mcbxv.2r5
++++++ warning: implicit element re-definition ignored: mqxb.b2r5
++++++ warning: implicit element re-definition ignored: tasb.3r5
++++++ warning: implicit element re-definition ignored: mqsx.3r5
++++++ warning: implicit element re-definition ignored: mqxa.3r5
++++++ warning: implicit element re-definition ignored: mcbxh.3r5
++++++ warning: implicit element re-definition ignored: mcbxv.3r5
++++++ warning: implicit element re-definition ignored: mcsx.3r5
++++++ warning: implicit element re-definition ignored: mctx.3r5
++++++ warning: implicit element re-definition ignored: mcosx.3r5
++++++ warning: implicit element re-definition ignored: mcox.3r5
++++++ warning: implicit element re-definition ignored: mcssx.3r5
++++++ warning: implicit element re-definition ignored: dfbxf.3r5
++++++ warning: implicit element re-definition ignored: x5fca.b4r5
++++++ warning: implicit element re-definition ignored: mbxw.a4r5
++++++ warning: implicit element re-definition ignored: mbxw.b4r5
++++++ warning: implicit element re-definition ignored: mbxw.c4r5
++++++ warning: implicit element re-definition ignored: mbxw.d4r5
++++++ warning: implicit element re-definition ignored: mbxw.e4r5
++++++ warning: implicit element re-definition ignored: mbxw.f4r5
++++++ warning: implicit element re-definition ignored: x5fca.a4r5
++++++ warning: implicit element re-definition ignored: x5fcb.a4r5
++++++ warning: implicit element re-definition ignored: x5zdc.b4r5
++++++ warning: implicit element re-definition ignored: brana.4r5
++++++ warning: implicit element re-definition ignored: x5zdc.a4r5
++++++ warning: implicit element re-definition ignored: tanc.4r5
++++++ warning: implicit element re-definition ignored: ip6
++++++ warning: implicit element re-definition ignored: ip7
++++++ warning: implicit element re-definition ignored: branb.4l8
++++++ warning: implicit element re-definition ignored: tclia.4l8
++++++ warning: implicit element re-definition ignored: mbx.4l8
++++++ warning: implicit element re-definition ignored: dfbxg.3l8
++++++ warning: implicit element re-definition ignored: mcosx.3l8
++++++ warning: implicit element re-definition ignored: mcox.3l8
++++++ warning: implicit element re-definition ignored: mcssx.3l8
++++++ warning: implicit element re-definition ignored: mcbxh.3l8
++++++ warning: implicit element re-definition ignored: mcbxv.3l8
++++++ warning: implicit element re-definition ignored: mcsx.3l8
++++++ warning: implicit element re-definition ignored: mctx.3l8
++++++ warning: implicit element re-definition ignored: mqxa.3l8
++++++ warning: implicit element re-definition ignored: mqsx.3l8
++++++ warning: implicit element re-definition ignored: mqxb.b2l8
++++++ warning: implicit element re-definition ignored: mcbxh.2l8
++++++ warning: implicit element re-definition ignored: mcbxv.2l8
++++++ warning: implicit element re-definition ignored: mqxb.a2l8
++++++ warning: implicit element re-definition ignored: mcbxh.1l8
++++++ warning: implicit element re-definition ignored: mcbxv.1l8
++++++ warning: implicit element re-definition ignored: mqxa.1l8
++++++ warning: implicit element re-definition ignored: mbxws.1l8
++++++ warning: implicit element re-definition ignored: mbxwh.1l8
++++++ warning: implicit element re-definition ignored: ip8
++++++ warning: implicit element re-definition ignored: mblw.1r8
++++++ warning: implicit element re-definition ignored: mbxws.1r8
++++++ warning: implicit element re-definition ignored: mqxa.1r8
++++++ warning: implicit element re-definition ignored: mcbxh.1r8
++++++ warning: implicit element re-definition ignored: mcbxv.1r8
++++++ warning: implicit element re-definition ignored: mqxb.a2r8
++++++ warning: implicit element re-definition ignored: mcbxh.2r8
++++++ warning: implicit element re-definition ignored: mcbxv.2r8
++++++ warning: implicit element re-definition ignored: mqxb.b2r8
++++++ warning: implicit element re-definition ignored: mqsx.3r8
++++++ warning: implicit element re-definition ignored: mqxa.3r8
++++++ warning: implicit element re-definition ignored: mcbxh.3r8
++++++ warning: implicit element re-definition ignored: mcbxv.3r8
++++++ warning: implicit element re-definition ignored: mcsx.3r8
++++++ warning: implicit element re-definition ignored: mctx.3r8
++++++ warning: implicit element re-definition ignored: mcosx.3r8
++++++ warning: implicit element re-definition ignored: mcox.3r8
++++++ warning: implicit element re-definition ignored: mcssx.3r8
++++++ warning: implicit element re-definition ignored: dfbxh.3r8
++++++ warning: implicit element re-definition ignored: mbx.4r8
++++++ warning: implicit element re-definition ignored: tcddm.4r8
++++++ warning: implicit element re-definition ignored: btvst.a4r8
++++++ warning: implicit element re-definition ignored: branb.4r8
++++++ warning: implicit element re-definition ignored: tanal.4l1
++++++ warning: implicit element re-definition ignored: x1zdc.a4l1
++++++ warning: implicit element re-definition ignored: brana.4l1
++++++ warning: implicit element re-definition ignored: x1fcl.4l1
++++++ warning: implicit element re-definition ignored: mbxw.f4l1
++++++ warning: implicit element re-definition ignored: mbxw.e4l1
++++++ warning: implicit element re-definition ignored: mbxw.d4l1
++++++ warning: implicit element re-definition ignored: mbxw.c4l1
++++++ warning: implicit element re-definition ignored: mbxw.b4l1
++++++ warning: implicit element re-definition ignored: mbxw.a4l1
++++++ warning: implicit element re-definition ignored: dfbxa.3l1
++++++ warning: implicit element re-definition ignored: mcosx.3l1
++++++ warning: implicit element re-definition ignored: mcox.3l1
++++++ warning: implicit element re-definition ignored: mcssx.3l1
++++++ warning: implicit element re-definition ignored: mcbxh.3l1
++++++ warning: implicit element re-definition ignored: mcbxv.3l1
++++++ warning: implicit element re-definition ignored: mcsx.3l1
++++++ warning: implicit element re-definition ignored: mctx.3l1
++++++ warning: implicit element re-definition ignored: mqxa.3l1
++++++ warning: implicit element re-definition ignored: mqsx.3l1
++++++ warning: implicit element re-definition ignored: tasb.3l1
++++++ warning: implicit element re-definition ignored: mqxb.b2l1
++++++ warning: implicit element re-definition ignored: mcbxh.2l1
++++++ warning: implicit element re-definition ignored: mcbxv.2l1
++++++ warning: implicit element re-definition ignored: mqxb.a2l1
++++++ warning: implicit element re-definition ignored: mcbxh.1l1
++++++ warning: implicit element re-definition ignored: mcbxv.1l1
++++++ warning: implicit element re-definition ignored: mqxa.1l1
++++++ warning: implicit element re-definition ignored: bpmwk.1l1
++++++ warning: implicit element re-definition ignored: tas.1l1
++++++ warning: implicit element re-definition ignored: mbas2.1l1
++++++ warning: implicit element re-definition ignored: ip1.l1
enter Twiss module
iteration: 1 error: 2.525108E-03 deltap: 0.000000E+00
orbit: -3.056898E-05 6.269177E-08 4.857955E-06 -1.448609E-04 0.000000E+00 0.000000E+00
iteration: 2 error: 4.315597E-05 deltap: 0.000000E+00
orbit: 1.134273E-09 -7.053445E-10 -2.408938E-10 -1.450000E-04 0.000000E+00 0.000000E+00
iteration: 3 error: 4.337533E-09 deltap: 0.000000E+00
orbit: 2.590299E-11 -5.470202E-10 1.672655E-13 -1.450000E-04 0.000000E+00 0.000000E+00
++++++ table: summ
length orbit5 alfa gammatr
2.665888319999896e+04 -0.000000000000000e+00 3.202048804239000e-04 5.588381245759998e+01
q1 dq1 betxmax dxmax
6.430999990172486e+01 3.000000078069948e+00 5.935115009636181e+02 2.720741306485160e+00
dxrms xcomax xcorms q2
1.344104534674799e+00 1.545439897281311e-02 9.885626081800364e-04 5.932000002666296e+01
dq2 betymax dymax dyrms
2.999999805244852e+00 6.098696059736556e+02 1.210194688464986e-01 3.643142661904077e-02
ycomax ycorms deltap synch_1
6.986494064697167e-03 6.273649315943959e-04 0.000000000000000e+00 0.000000000000000e+00
synch_2 synch_3 synch_4 synch_5
0.000000000000000e+00 0.000000000000000e+00 0.000000000000000e+00 0.000000000000000e+00
nflips
0.000000000000000e+00
enter Twiss module
iteration: 1 error: 2.517161E-03 deltap: 0.000000E+00
orbit: 1.769414E-05 -5.593396E-07 -2.801367E-06 1.450025E-04 0.000000E+00 0.000000E+00
iteration: 2 error: 3.438873E-05 deltap: 0.000000E+00
orbit: 2.365727E-09 3.662869E-10 -1.633338E-10 1.450000E-04 0.000000E+00 0.000000E+00
iteration: 3 error: 5.623722E-10 deltap: 0.000000E+00
orbit: 1.323483E-09 3.178942E-10 1.104578E-13 1.450000E-04 0.000000E+00 0.000000E+00
++++++ table: summ
length orbit5 alfa gammatr
2.665888319999895e+04 -0.000000000000000e+00 3.192054999491873e-04 5.597122565324666e+01
q1 dq1 betxmax dxmax
6.430999990601576e+01 3.000000009898441e+00 5.909961120775461e+02 2.921792801755544e+00
dxrms xcomax xcorms q2
1.343382960241845e+00 1.545439614740161e-02 9.994160708666150e-04 5.932000002847266e+01
dq2 betymax dymax dyrms
3.000000010147829e+00 6.046420530599705e+02 1.272431476595715e-01 2.218671992885489e-02
ycomax ycorms deltap synch_1
6.986494063941867e-03 6.278667310642880e-04 0.000000000000000e+00 0.000000000000000e+00
synch_2 synch_3 synch_4 synch_5
0.000000000000000e+00 0.000000000000000e+00 0.000000000000000e+00 0.000000000000000e+00
nflips
0.000000000000000e+00
True
list(myMad.table)
['summ', 'twiss_ipb1_flat', 'twiss_ipb2_flat']
if 1:
b1DF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/b1DF.pickle')
else:
b1DF=myMad.table.twiss_ipb1_flat.dframe()
b1DF.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/b1DF.pickle')
if 1:
b2DF=pd.read_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/b2DF.pickle')
else:
b2DF=myMad.table.twiss_ipb2_flat.dframe()
b2DF.to_pickle('/eos/user/s/sterbini/MD_ANALYSIS/2019/VdMScans/b2DF.pickle')